Probe CUST_17414_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17414_PI426222305 | JHI_St_60k_v1 | DMT400001518 | CAGTTTCAATGGGATCCTGTCGAAGGAGATGATGTTGATCTCTCGGAGAAGCTAATATTC |
All Microarray Probes Designed to Gene DMG400000564
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17414_PI426222305 | JHI_St_60k_v1 | DMT400001518 | CAGTTTCAATGGGATCCTGTCGAAGGAGATGATGTTGATCTCTCGGAGAAGCTAATATTC |
Microarray Signals from CUST_17414_PI426222305
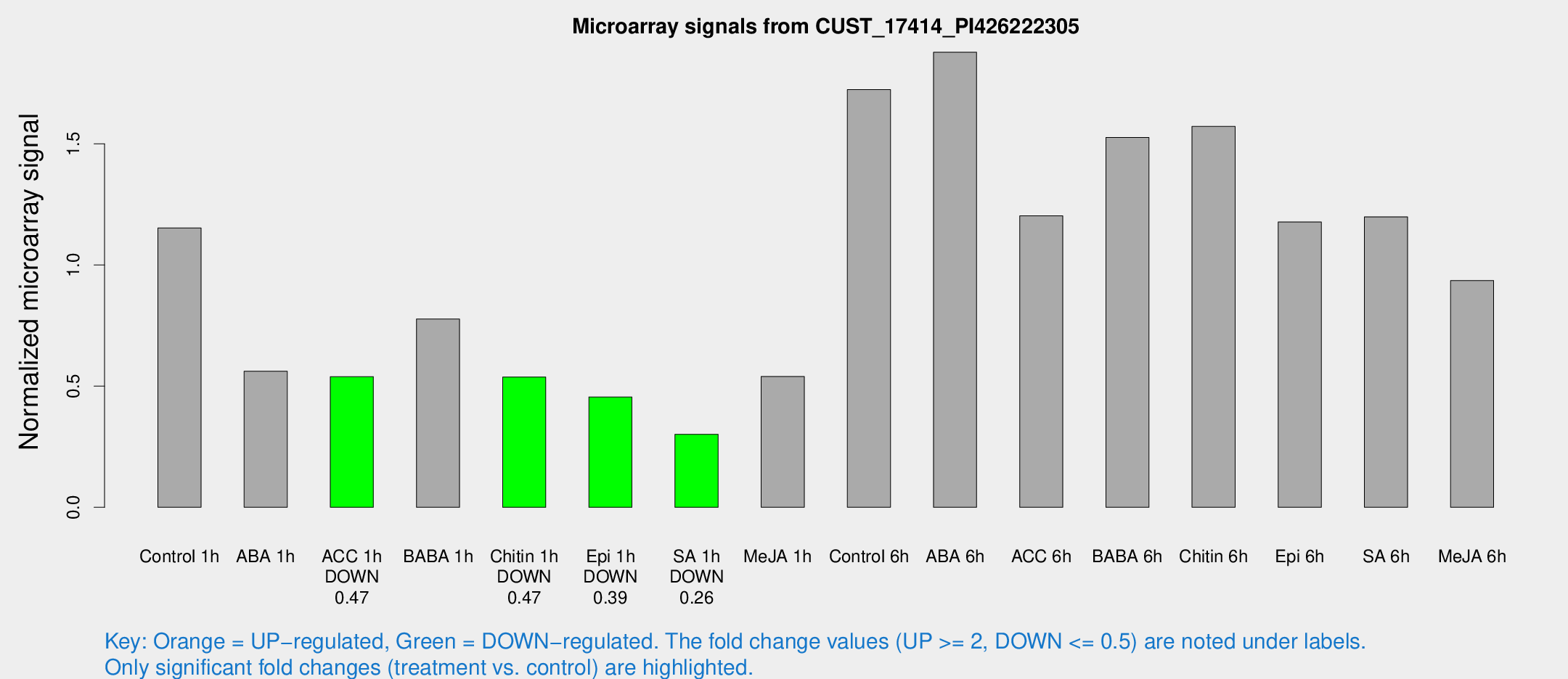
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 19.864 | 3.21711 | 1.15309 | 0.19178 |
| ABA 1h | 10.819 | 5.65129 | 0.561627 | 0.271736 |
| ACC 1h | 9.832 | 3.34854 | 0.539108 | 0.209807 |
| BABA 1h | 12.8414 | 3.26158 | 0.776957 | 0.200913 |
| Chitin 1h | 8.52315 | 3.10021 | 0.537631 | 0.205241 |
| Epi 1h | 6.77064 | 3.01681 | 0.455196 | 0.206396 |
| SA 1h | 5.2386 | 3.01434 | 0.300944 | 0.17302 |
| Me-JA 1h | 8.22359 | 3.14855 | 0.539582 | 0.254162 |
| Control 6h | 30.0227 | 4.90346 | 1.72375 | 0.221448 |
| ABA 6h | 33.8162 | 3.8175 | 1.8781 | 0.211997 |
| ACC 6h | 25.251 | 6.54304 | 1.20285 | 0.231139 |
| BABA 6h | 28.9946 | 3.94255 | 1.52648 | 0.209792 |
| Chitin 6h | 28.7188 | 3.87067 | 1.57189 | 0.217531 |
| Epi 6h | 23.2726 | 4.12978 | 1.17756 | 0.328114 |
| SA 6h | 27.2782 | 11.729 | 1.19868 | 0.750988 |
| Me-JA 6h | 16.4986 | 3.59787 | 0.935729 | 0.207478 |
Source Transcript PGSC0003DMT400001518 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G64950.1 | +1 | 4e-179 | 518 | 283/524 (54%) | cytochrome P450, family 89, subfamily A, polypeptide 5 | chr1:24127587-24129119 FORWARD LENGTH=510 |
