Probe CUST_17388_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17388_PI426222305 | JHI_St_60k_v1 | DMT400001571 | AGTATACATCCGCCAAATTAAACTAATCGCGTTGGGTCTATTGTACACCATATATATTTG |
All Microarray Probes Designed to Gene DMG400000583
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17388_PI426222305 | JHI_St_60k_v1 | DMT400001571 | AGTATACATCCGCCAAATTAAACTAATCGCGTTGGGTCTATTGTACACCATATATATTTG |
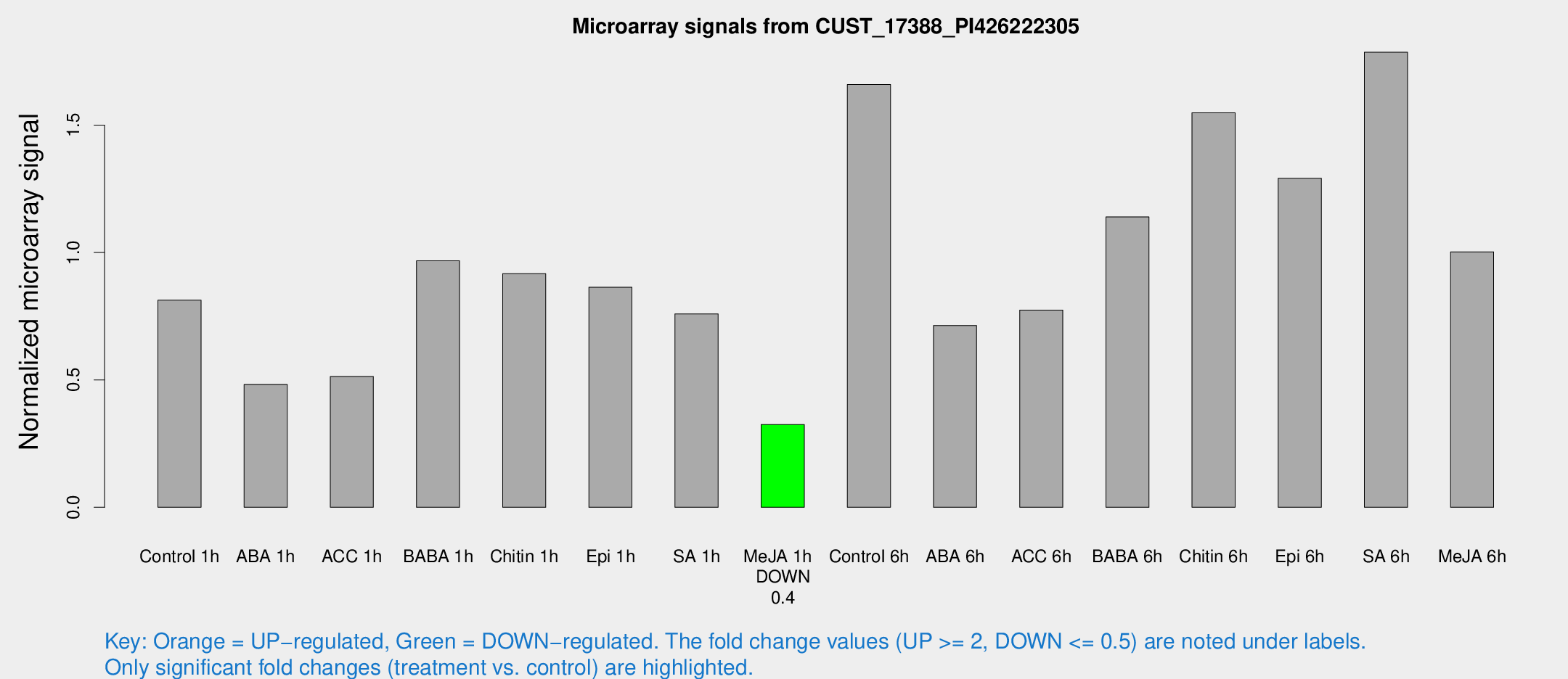
Microarray Signals from CUST_17388_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 57.6393 | 10.4568 | 0.812632 | 0.0934623 |
| ABA 1h | 35.1734 | 14.7018 | 0.48216 | 0.225344 |
| ACC 1h | 46.8687 | 18.6209 | 0.513288 | 0.30875 |
| BABA 1h | 69.3869 | 17.3192 | 0.967357 | 0.194884 |
| Chitin 1h | 60.6078 | 16.6837 | 0.9169 | 0.177714 |
| Epi 1h | 51.9674 | 5.91627 | 0.863513 | 0.0804609 |
| SA 1h | 54.4638 | 7.1935 | 0.758568 | 0.0641669 |
| Me-JA 1h | 18.2891 | 3.48159 | 0.325055 | 0.0619272 |
| Control 6h | 117.658 | 19.0208 | 1.65914 | 0.145268 |
| ABA 6h | 52.5704 | 4.89101 | 0.713656 | 0.0748136 |
| ACC 6h | 64.0096 | 13.188 | 0.773947 | 0.170927 |
| BABA 6h | 88.2391 | 7.9492 | 1.13941 | 0.0943733 |
| Chitin 6h | 114.348 | 12.2693 | 1.54787 | 0.120273 |
| Epi 6h | 100.055 | 7.02252 | 1.29119 | 0.0904173 |
| SA 6h | 121.694 | 7.91996 | 1.78583 | 0.160554 |
| Me-JA 6h | 74.555 | 20.6375 | 1.00245 | 0.21963 |
Source Transcript PGSC0003DMT400001571 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G53730.1 | +3 | 6e-71 | 221 | 124/215 (58%) | Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family | chr5:21808072-21808713 REVERSE LENGTH=213 |
