Probe CUST_17243_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17243_PI426222305 | JHI_St_60k_v1 | DMT400013822 | TGACTATCATTTTCCTGCTCCTTGGGTAGGACCATTGAAGAATGCAATTGATTGGGCCTC |
All Microarray Probes Designed to Gene DMG400005404
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17326_PI426222305 | JHI_St_60k_v1 | DMT400013821 | GCTGATTGTTTCCTTTTTGCTTCTCCAGAGTATAATTACTCCATCACAGACTAAATAGAA |
| CUST_17243_PI426222305 | JHI_St_60k_v1 | DMT400013822 | TGACTATCATTTTCCTGCTCCTTGGGTAGGACCATTGAAGAATGCAATTGATTGGGCCTC |
Microarray Signals from CUST_17243_PI426222305
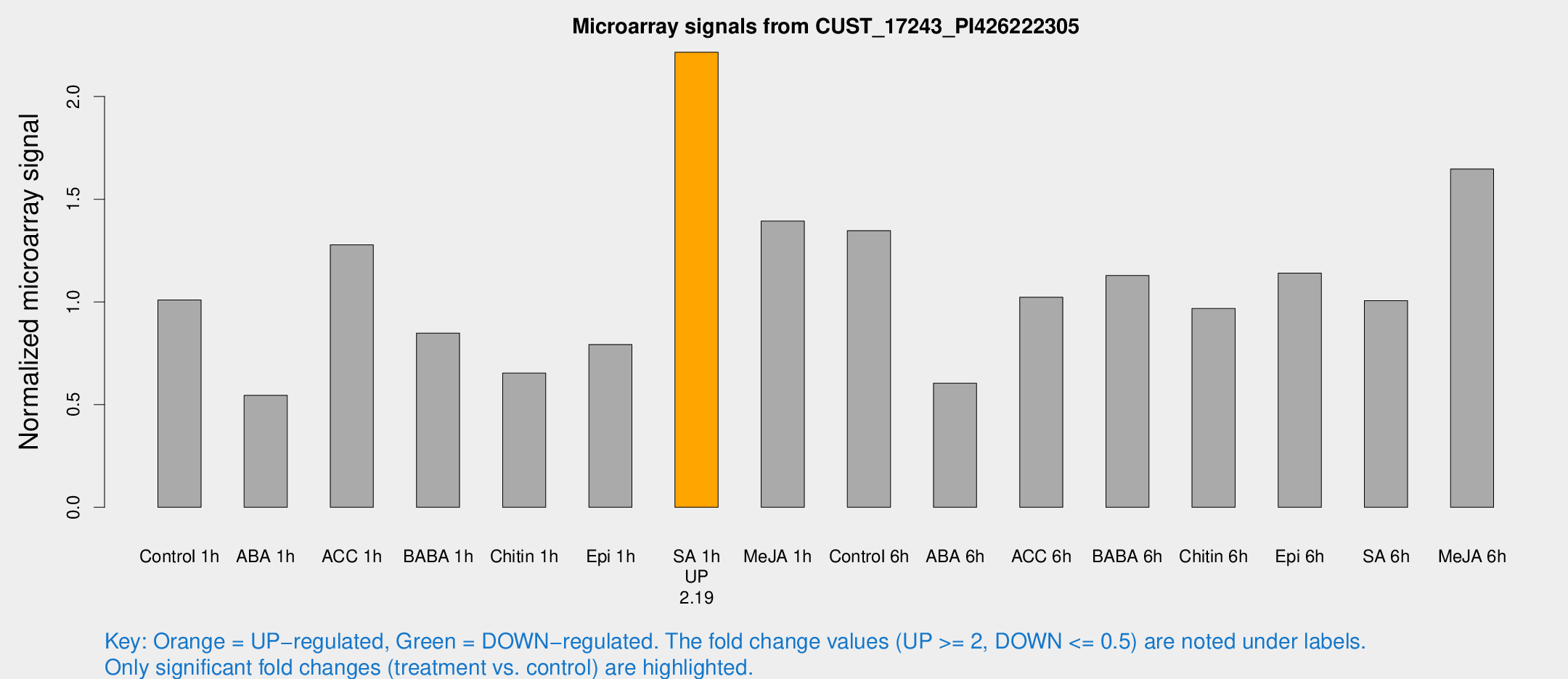
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 245.563 | 43.9263 | 1.00996 | 0.109788 |
| ABA 1h | 117.9 | 21.6582 | 0.544925 | 0.0910777 |
| ACC 1h | 356.192 | 111.411 | 1.27794 | 0.458523 |
| BABA 1h | 196.29 | 26.067 | 0.848218 | 0.0542561 |
| Chitin 1h | 138.881 | 8.75003 | 0.653608 | 0.0411046 |
| Epi 1h | 163.73 | 17.7564 | 0.792633 | 0.0857172 |
| SA 1h | 562.408 | 111.517 | 2.21589 | 0.406181 |
| Me-JA 1h | 274.297 | 42.3432 | 1.39356 | 0.121834 |
| Control 6h | 349.527 | 106.494 | 1.34736 | 0.31084 |
| ABA 6h | 154.49 | 21.8776 | 0.604072 | 0.0515421 |
| ACC 6h | 279.548 | 27.4377 | 1.02297 | 0.109599 |
| BABA 6h | 299.062 | 18.6569 | 1.12856 | 0.0669854 |
| Chitin 6h | 244.552 | 18.9189 | 0.968017 | 0.107939 |
| Epi 6h | 314.49 | 57.6838 | 1.13991 | 0.296371 |
| SA 6h | 245.567 | 48.789 | 1.00607 | 0.107359 |
| Me-JA 6h | 397.21 | 64.7987 | 1.64752 | 0.131597 |
Source Transcript PGSC0003DMT400013822 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G27890.1 | +3 | 9e-34 | 124 | 67/88 (76%) | NADPH:quinone oxidoreductase | chr3:10350807-10351938 REVERSE LENGTH=196 |
