Probe CUST_17137_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17137_PI426222305 | JHI_St_60k_v1 | DMT400018339 | CCATGGTGTCCATCTAGGGAAGAGTAACAATTCTTTTTCCTACTCAAAATATGAGACGTG |
All Microarray Probes Designed to Gene DMG400007122
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17137_PI426222305 | JHI_St_60k_v1 | DMT400018339 | CCATGGTGTCCATCTAGGGAAGAGTAACAATTCTTTTTCCTACTCAAAATATGAGACGTG |
Microarray Signals from CUST_17137_PI426222305
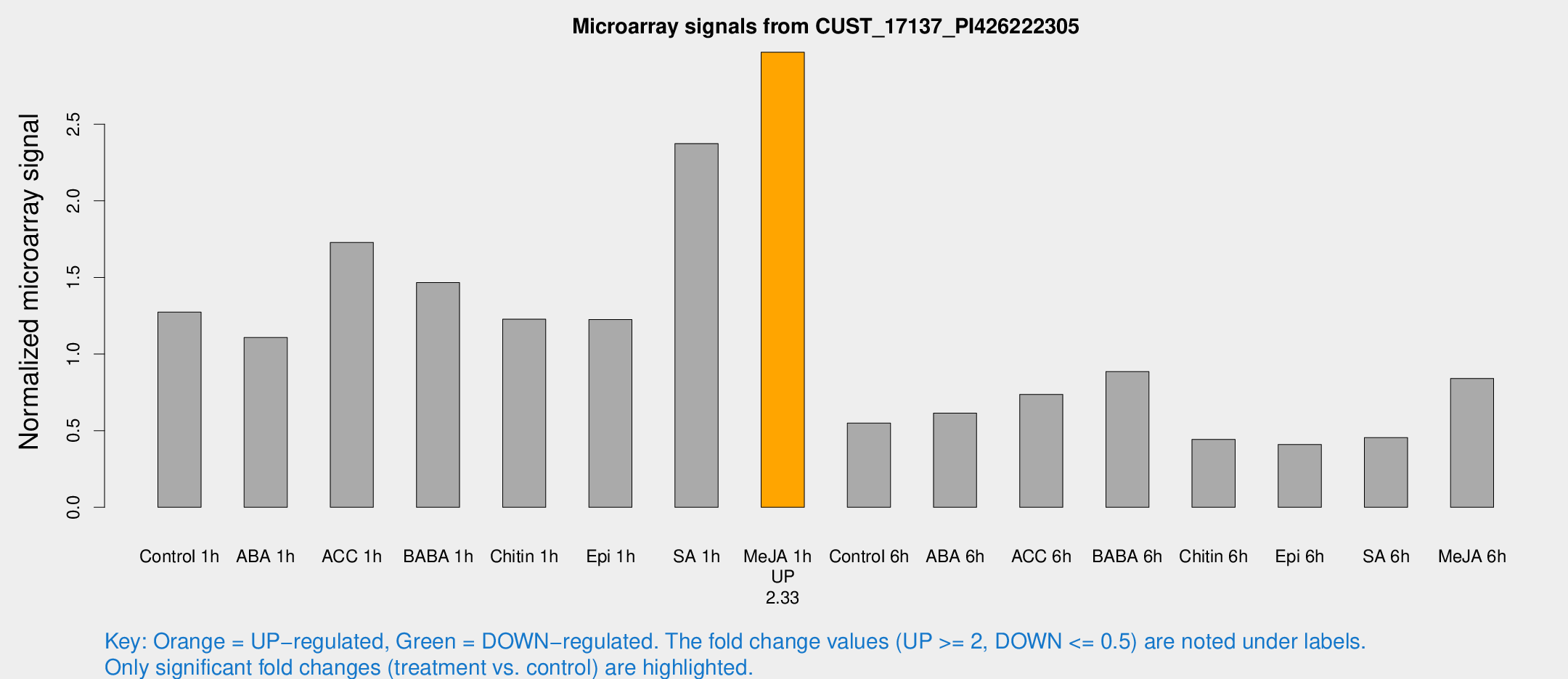
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8990.4 | 1776 | 1.27387 | 0.16568 |
| ABA 1h | 6802.39 | 976.57 | 1.10771 | 0.0836035 |
| ACC 1h | 13570.8 | 4050.07 | 1.72843 | 0.530217 |
| BABA 1h | 10038.8 | 1882.19 | 1.46645 | 0.158183 |
| Chitin 1h | 7585.38 | 730.013 | 1.2274 | 0.0708664 |
| Epi 1h | 7230.09 | 417.94 | 1.22518 | 0.0707382 |
| SA 1h | 18046.6 | 5182.51 | 2.37359 | 0.664404 |
| Me-JA 1h | 16686.7 | 1664.61 | 2.96957 | 0.17145 |
| Control 6h | 4072.26 | 1118.88 | 0.549568 | 0.114679 |
| ABA 6h | 4809.35 | 1302.97 | 0.614007 | 0.147186 |
| ACC 6h | 5901.18 | 870.665 | 0.736994 | 0.118271 |
| BABA 6h | 6973.32 | 1283.87 | 0.885863 | 0.143472 |
| Chitin 6h | 3216.14 | 185.859 | 0.44335 | 0.0256026 |
| Epi 6h | 3155.32 | 182.8 | 0.409645 | 0.0449158 |
| SA 6h | 3067.07 | 177.501 | 0.454317 | 0.0349983 |
| Me-JA 6h | 5840.49 | 925.548 | 0.840656 | 0.109333 |
Source Transcript PGSC0003DMT400018339 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G27820.1 | +1 | 0.0 | 586 | 306/379 (81%) | prephenate dehydratase 1 | chr2:11856808-11858082 FORWARD LENGTH=424 |
