Probe CUST_16815_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16815_PI426222305 | JHI_St_60k_v1 | DMT400069394 | CTTGCCTACTTACAAATACTGTTAAGGTTGTTCTTGAGAGAAAATGATCAAAATGGTCCT |
All Microarray Probes Designed to Gene DMG401026984
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16815_PI426222305 | JHI_St_60k_v1 | DMT400069394 | CTTGCCTACTTACAAATACTGTTAAGGTTGTTCTTGAGAGAAAATGATCAAAATGGTCCT |
| CUST_16591_PI426222305 | JHI_St_60k_v1 | DMT400069392 | CGATTAGGCAGAATCAATCTTTCATAAGTCCTACTGTAAAGGTTTTGTACTGTGTTAATC |
Microarray Signals from CUST_16815_PI426222305
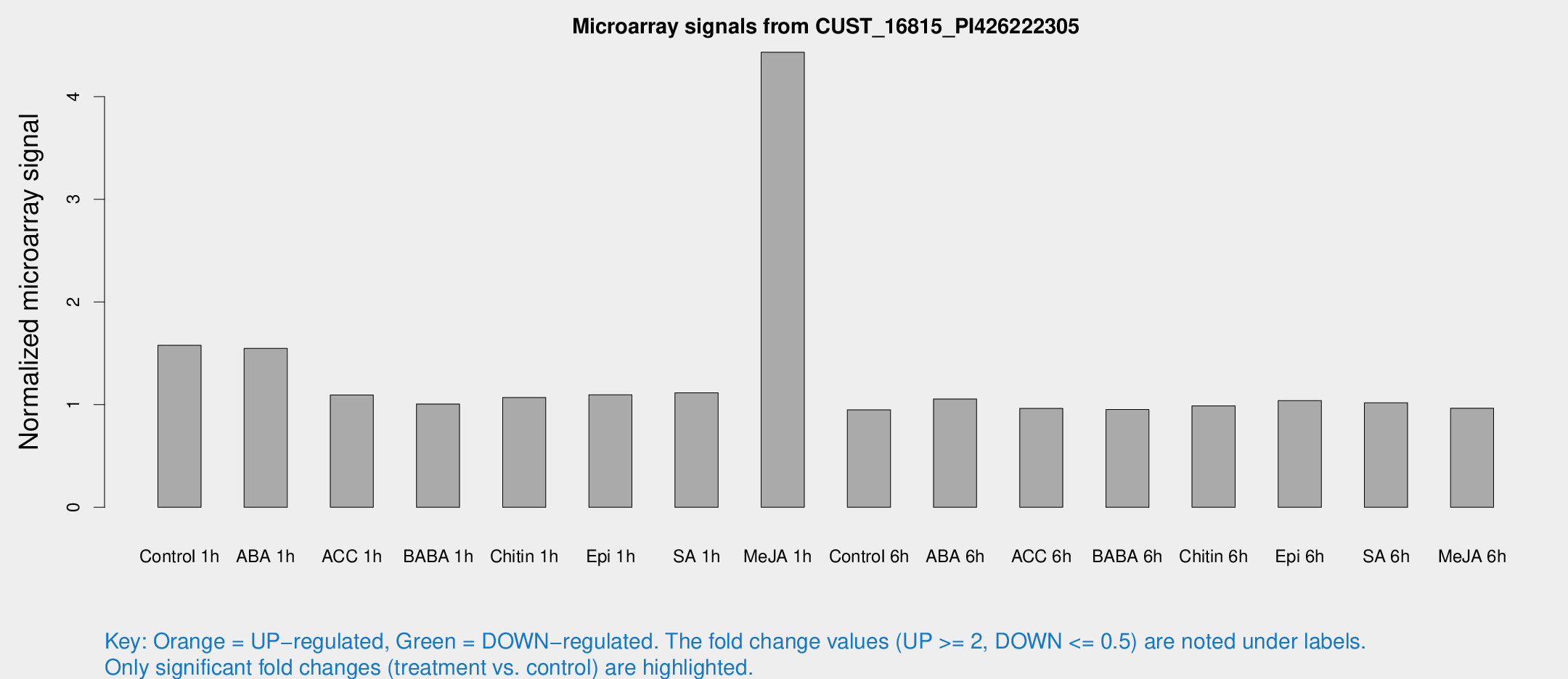
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8.82344 | 2.88313 | 1.57718 | 0.577457 |
| ABA 1h | 7.5453 | 2.76451 | 1.5482 | 0.61807 |
| ACC 1h | 6.07491 | 3.1115 | 1.09419 | 0.572675 |
| BABA 1h | 5.18075 | 3.00361 | 1.00634 | 0.580905 |
| Chitin 1h | 4.90623 | 2.85603 | 1.0685 | 0.589529 |
| Epi 1h | 4.90546 | 2.8486 | 1.09555 | 0.613212 |
| SA 1h | 6.37898 | 2.8745 | 1.1148 | 0.547294 |
| Me-JA 1h | 19.4275 | 3.11885 | 4.43267 | 0.712266 |
| Control 6h | 5.07588 | 2.94042 | 0.948515 | 0.547763 |
| ABA 6h | 6.05887 | 3.0808 | 1.05516 | 0.549191 |
| ACC 6h | 6.03919 | 3.57658 | 0.962465 | 0.557335 |
| BABA 6h | 5.71951 | 3.31886 | 0.952375 | 0.551502 |
| Chitin 6h | 5.63631 | 3.26634 | 0.988712 | 0.572876 |
| Epi 6h | 6.45783 | 3.41163 | 1.03929 | 0.559729 |
| SA 6h | 5.3909 | 3.12381 | 1.01722 | 0.589052 |
| Me-JA 6h | 5.08314 | 2.94707 | 0.965284 | 0.551914 |
Source Transcript PGSC0003DMT400069394 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G63160.1 | +3 | 3e-125 | 370 | 186/293 (63%) | BTB and TAZ domain protein 1 | chr5:25333485-25335399 REVERSE LENGTH=365 |
