Probe CUST_16659_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16659_PI426222305 | JHI_St_60k_v1 | DMT400069546 | CACGATTTACGCTAGTTTGTTATTCGAATTTGAGCAGGTCATTATGTATCAACTATGGTA |
All Microarray Probes Designed to Gene DMG400027035
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16593_PI426222305 | JHI_St_60k_v1 | DMT400069547 | CACGATTTACGCTAGTTTGTTATTCGAATTTGAGCAGGTCATTATGTATCAACTATGGTA |
| CUST_16659_PI426222305 | JHI_St_60k_v1 | DMT400069546 | CACGATTTACGCTAGTTTGTTATTCGAATTTGAGCAGGTCATTATGTATCAACTATGGTA |
Microarray Signals from CUST_16659_PI426222305
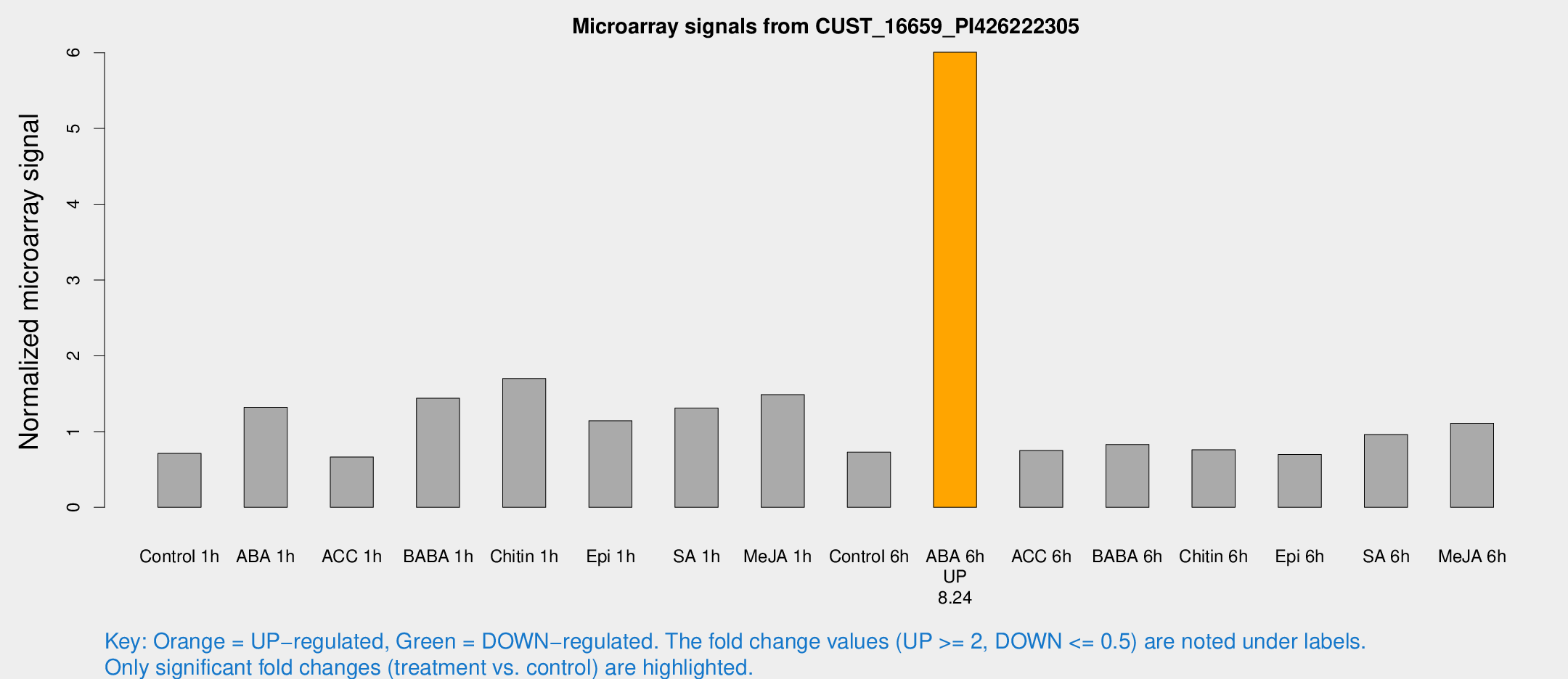
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 237.432 | 29.2862 | 0.712619 | 0.0837695 |
| ABA 1h | 398.752 | 76.7428 | 1.31885 | 0.233233 |
| ACC 1h | 244.076 | 69.1191 | 0.663064 | 0.16693 |
| BABA 1h | 482.921 | 110.191 | 1.43952 | 0.226699 |
| Chitin 1h | 534.025 | 131.342 | 1.69946 | 0.403729 |
| Epi 1h | 334.262 | 54.6037 | 1.14353 | 0.194574 |
| SA 1h | 456.192 | 86.4489 | 1.31012 | 0.173539 |
| Me-JA 1h | 401.195 | 34.8045 | 1.48638 | 0.0865287 |
| Control 6h | 255.751 | 60.6126 | 0.729311 | 0.135901 |
| ABA 6h | 2178.53 | 416.281 | 6.00615 | 0.905811 |
| ACC 6h | 292.256 | 51.8818 | 0.748966 | 0.101211 |
| BABA 6h | 308.879 | 34.7477 | 0.829886 | 0.0715032 |
| Chitin 6h | 265.644 | 15.7326 | 0.758497 | 0.0543007 |
| Epi 6h | 262.865 | 35.0864 | 0.698363 | 0.0413627 |
| SA 6h | 316.292 | 37.133 | 0.960293 | 0.056363 |
| Me-JA 6h | 364.154 | 25.2822 | 1.10953 | 0.0647059 |
Source Transcript PGSC0003DMT400069546 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G16840.2 | +3 | 3e-56 | 195 | 118/207 (57%) | binding partner of acd11 1 | chr5:5536042-5538026 FORWARD LENGTH=260 |
