Probe CUST_16618_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16618_PI426222305 | JHI_St_60k_v1 | DMT400069145 | CCAAGACCTTGCTCGTTTTGCTGTTCAGGATTATAACAAGAAAGAGGTTAATTATAATGA |
All Microarray Probes Designed to Gene DMG400026899
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16618_PI426222305 | JHI_St_60k_v1 | DMT400069145 | CCAAGACCTTGCTCGTTTTGCTGTTCAGGATTATAACAAGAAAGAGGTTAATTATAATGA |
| CUST_16650_PI426222305 | JHI_St_60k_v1 | DMT400069147 | GGAGAATTTCAAGGAAGTTCAAGAATTCAAGCTTGTTGGTGATGCTACAAAGTAAAATGA |
Microarray Signals from CUST_16618_PI426222305
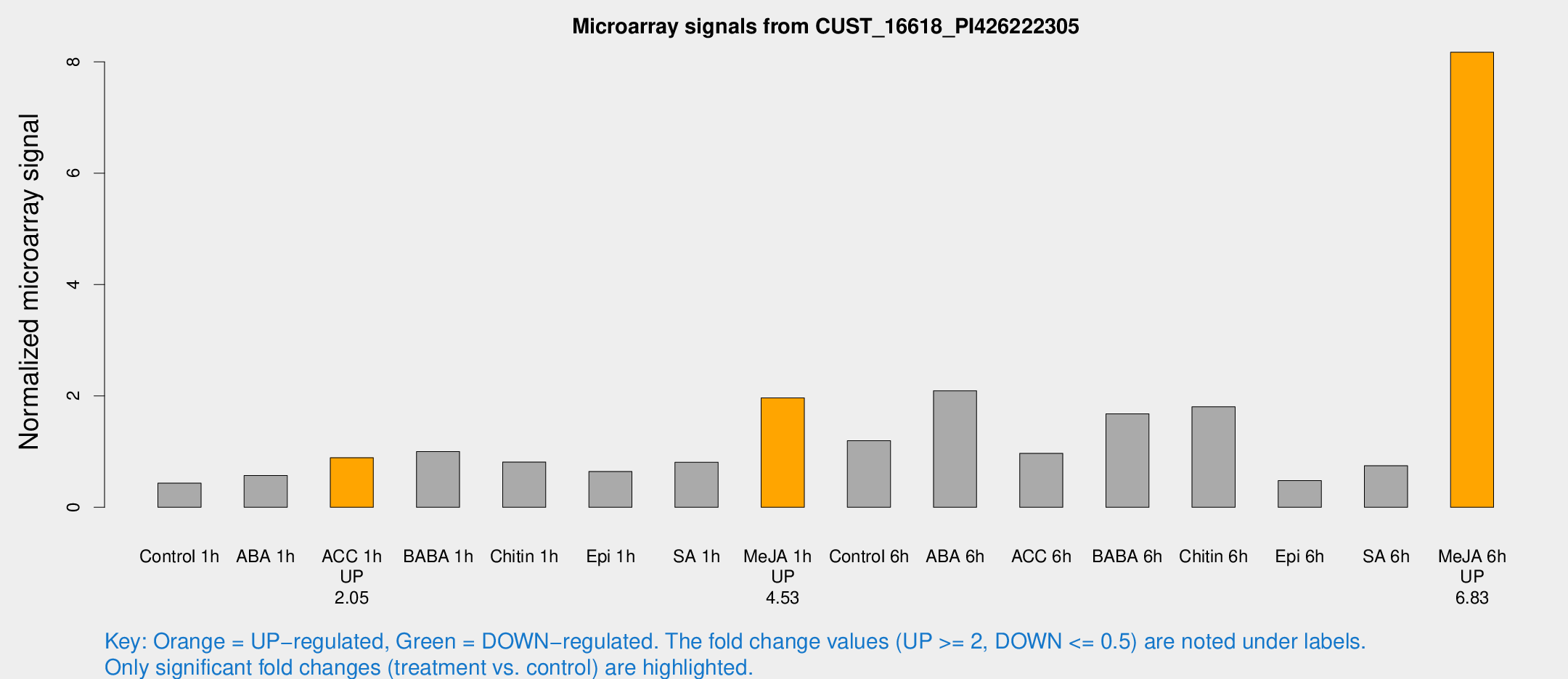
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2693.87 | 376.776 | 0.433921 | 0.0873625 |
| ABA 1h | 3634.2 | 1345.8 | 0.572285 | 0.227186 |
| ACC 1h | 5691.48 | 886.222 | 0.890394 | 0.132478 |
| BABA 1h | 5938.5 | 607.151 | 1.00166 | 0.0578335 |
| Chitin 1h | 4540.85 | 626.515 | 0.813081 | 0.0642844 |
| Epi 1h | 3595.99 | 887.605 | 0.643307 | 0.164087 |
| SA 1h | 5733.55 | 2099.48 | 0.810302 | 0.294424 |
| Me-JA 1h | 9841.51 | 920.94 | 1.9639 | 0.113388 |
| Control 6h | 9531.41 | 4671.77 | 1.19603 | 0.620329 |
| ABA 6h | 13596.1 | 925.187 | 2.09319 | 0.223641 |
| ACC 6h | 7086.7 | 1570.83 | 0.969415 | 0.107412 |
| BABA 6h | 11568.8 | 1281.4 | 1.6799 | 0.179943 |
| Chitin 6h | 12278.8 | 2822.36 | 1.80592 | 0.426689 |
| Epi 6h | 3777.98 | 1241.55 | 0.480164 | 0.266176 |
| SA 6h | 4688.46 | 1026.58 | 0.745863 | 0.126259 |
| Me-JA 6h | 52553.3 | 11783.4 | 8.17301 | 1.43662 |
Source Transcript PGSC0003DMT400069145 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G12490.2 | +2 | 6e-24 | 100 | 49/99 (49%) | cystatin B | chr3:3960523-3961876 REVERSE LENGTH=234 |
