Probe CUST_16433_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16433_PI426222305 | JHI_St_60k_v1 | DMT400027952 | AATTAATATTGTAACCAACCCCTACCCTCCTCTCCAGATACTAGTAGCAGTTATGCTCTG |
All Microarray Probes Designed to Gene DMG400010768
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16433_PI426222305 | JHI_St_60k_v1 | DMT400027952 | AATTAATATTGTAACCAACCCCTACCCTCCTCTCCAGATACTAGTAGCAGTTATGCTCTG |
Microarray Signals from CUST_16433_PI426222305
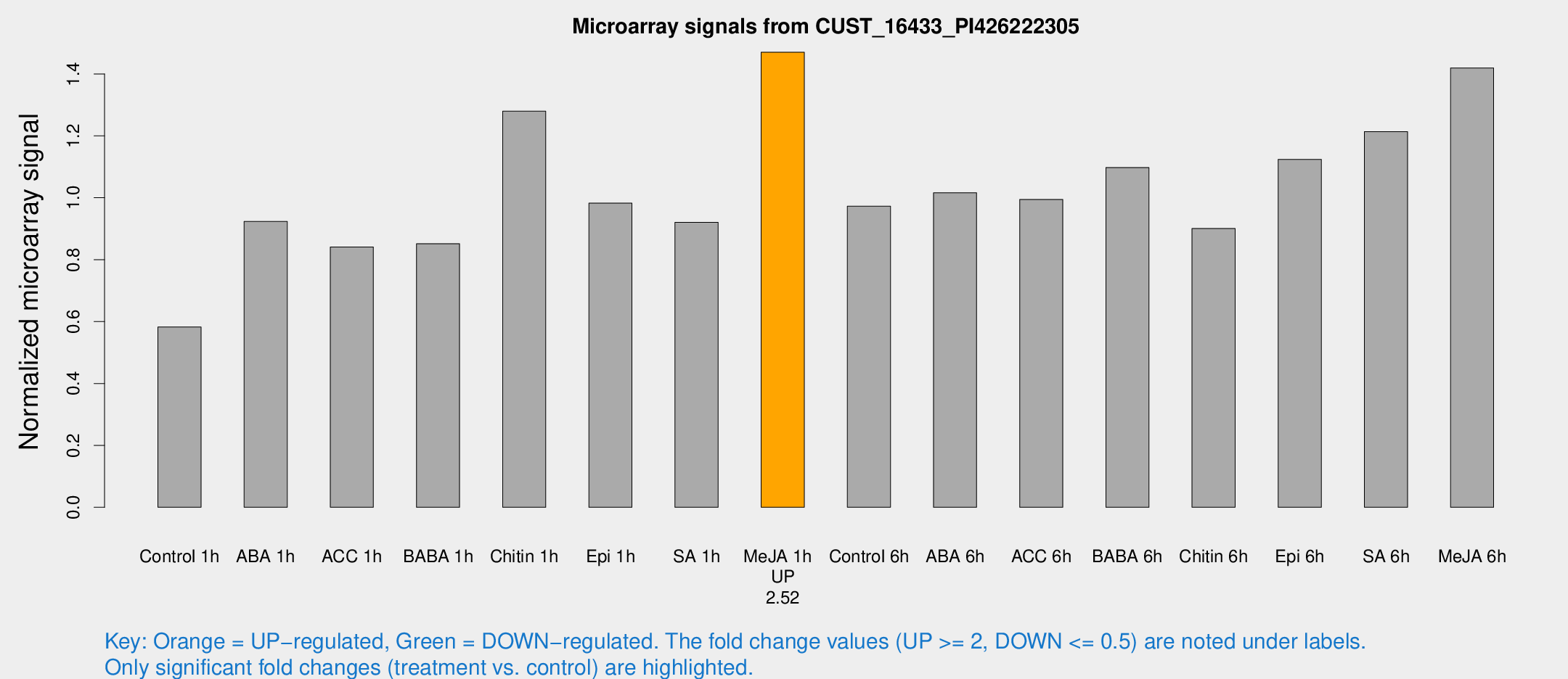
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 15.2369 | 3.52418 | 0.582751 | 0.137346 |
| ABA 1h | 23.1307 | 6.15098 | 0.92327 | 0.330529 |
| ACC 1h | 23.0595 | 3.94548 | 0.840741 | 0.182791 |
| BABA 1h | 24.4948 | 9.00737 | 0.851532 | 0.263547 |
| Chitin 1h | 31.1167 | 5.83685 | 1.27968 | 0.385043 |
| Epi 1h | 22.3529 | 3.66225 | 0.982556 | 0.1703 |
| SA 1h | 24.6613 | 3.721 | 0.920662 | 0.139483 |
| Me-JA 1h | 31.6045 | 3.9852 | 1.47016 | 0.188705 |
| Control 6h | 25.5035 | 3.91763 | 0.972591 | 0.151555 |
| ABA 6h | 28.2108 | 4.08639 | 1.01625 | 0.149495 |
| ACC 6h | 29.9428 | 4.77895 | 0.994699 | 0.154986 |
| BABA 6h | 32.1237 | 4.44773 | 1.09762 | 0.156121 |
| Chitin 6h | 26.7121 | 6.70911 | 0.900754 | 0.24444 |
| Epi 6h | 34.0348 | 5.88094 | 1.12374 | 0.158782 |
| SA 6h | 37.0456 | 15.1021 | 1.21384 | 0.58744 |
| Me-JA 6h | 37.8136 | 6.331 | 1.41938 | 0.329618 |
Source Transcript PGSC0003DMT400027952 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G03020.1 | +1 | 2e-32 | 115 | 55/94 (59%) | Thioredoxin superfamily protein | chr1:698207-698515 REVERSE LENGTH=102 |
