Probe CUST_16425_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16425_PI426222305 | JHI_St_60k_v1 | DMT400060584 | TCTTTGCCATGTGTCAAACGTGATATACATTTACCCAGTTCATGATACGTTCCTAACTTG |
All Microarray Probes Designed to Gene DMG400023566
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16425_PI426222305 | JHI_St_60k_v1 | DMT400060584 | TCTTTGCCATGTGTCAAACGTGATATACATTTACCCAGTTCATGATACGTTCCTAACTTG |
| CUST_16459_PI426222305 | JHI_St_60k_v1 | DMT400060583 | TAAATCCGGAGATCAGGTTCCACTTTATGCTAACAAAGTCGGTCCTTTCCATAATCCGAG |
Microarray Signals from CUST_16425_PI426222305
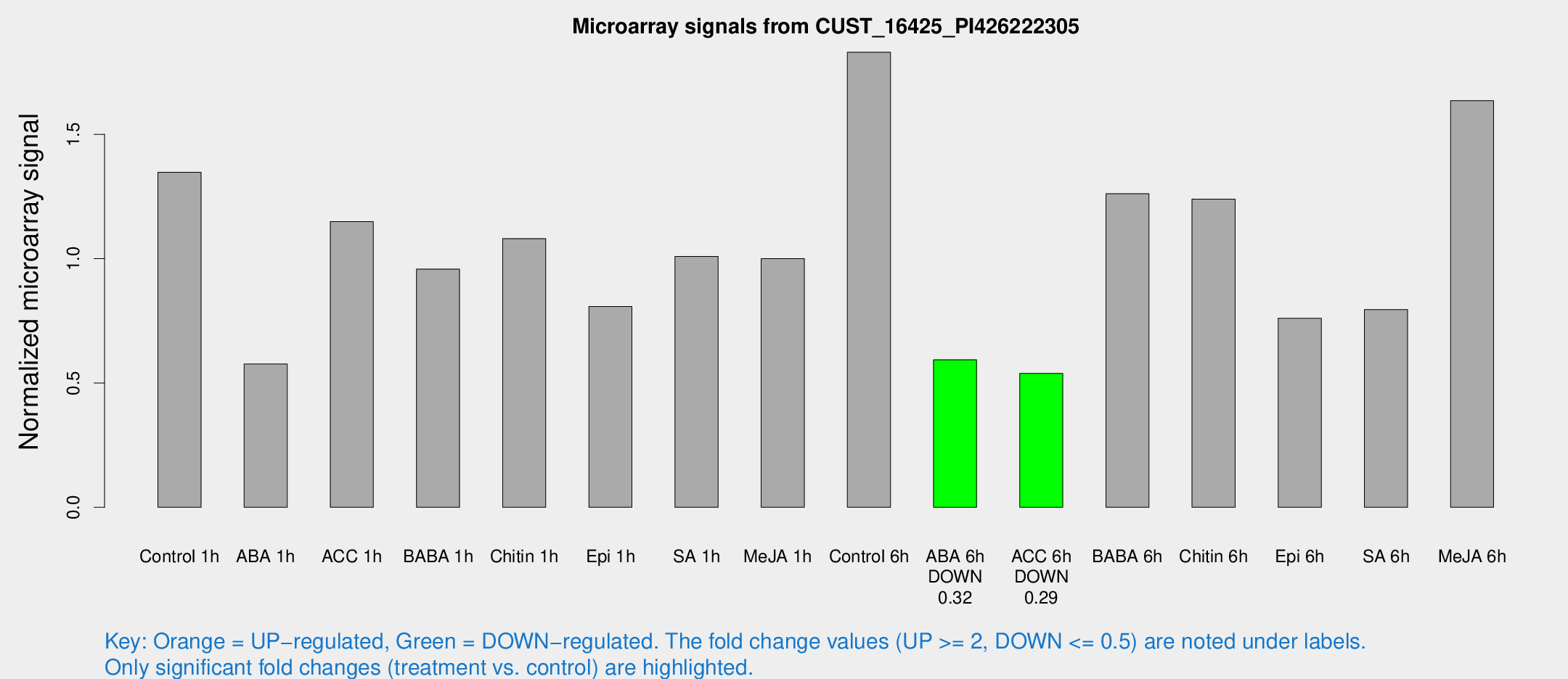
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 15.7058 | 3.02047 | 1.34704 | 0.26899 |
| ABA 1h | 5.8484 | 2.81251 | 0.576123 | 0.281207 |
| ACC 1h | 15.8351 | 5.30341 | 1.14865 | 0.440072 |
| BABA 1h | 12.5397 | 4.88777 | 0.957998 | 0.34659 |
| Chitin 1h | 11.5606 | 2.99536 | 1.08073 | 0.294847 |
| Epi 1h | 8.36441 | 2.91615 | 0.807522 | 0.316509 |
| SA 1h | 13.0383 | 3.41657 | 1.0091 | 0.423584 |
| Me-JA 1h | 10.6676 | 3.9461 | 0.99995 | 0.360321 |
| Control 6h | 21.5375 | 3.28035 | 1.83024 | 0.296722 |
| ABA 6h | 7.47583 | 3.16116 | 0.593436 | 0.265915 |
| ACC 6h | 7.35069 | 3.6887 | 0.538575 | 0.270258 |
| BABA 6h | 16.6763 | 3.54405 | 1.26109 | 0.29637 |
| Chitin 6h | 16.9497 | 5.81129 | 1.23931 | 0.434662 |
| Epi 6h | 11.0171 | 3.53933 | 0.760057 | 0.303935 |
| SA 6h | 10.7642 | 4.72679 | 0.795105 | 0.344717 |
| Me-JA 6h | 19.0462 | 3.23267 | 1.63578 | 0.28715 |
Source Transcript PGSC0003DMT400060584 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G14670.1 | +1 | 1e-83 | 276 | 150/162 (93%) | Endomembrane protein 70 protein family | chr1:5037669-5040199 FORWARD LENGTH=592 |
