Probe CUST_15922_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15922_PI426222305 | JHI_St_60k_v1 | DMT400076378 | AATCGATTTGCTGCATCTTCTATCTTCTGCTCTGAATTCCAAGTTTCTTGAATTCTTGAG |
All Microarray Probes Designed to Gene DMG400029702
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15922_PI426222305 | JHI_St_60k_v1 | DMT400076378 | AATCGATTTGCTGCATCTTCTATCTTCTGCTCTGAATTCCAAGTTTCTTGAATTCTTGAG |
| CUST_15980_PI426222305 | JHI_St_60k_v1 | DMT400076379 | CGGTTGTTGTTGGATTATAATGGGTATGTTGTTTGTTGAGCATACTGACTTTGTAATAGA |
Microarray Signals from CUST_15922_PI426222305
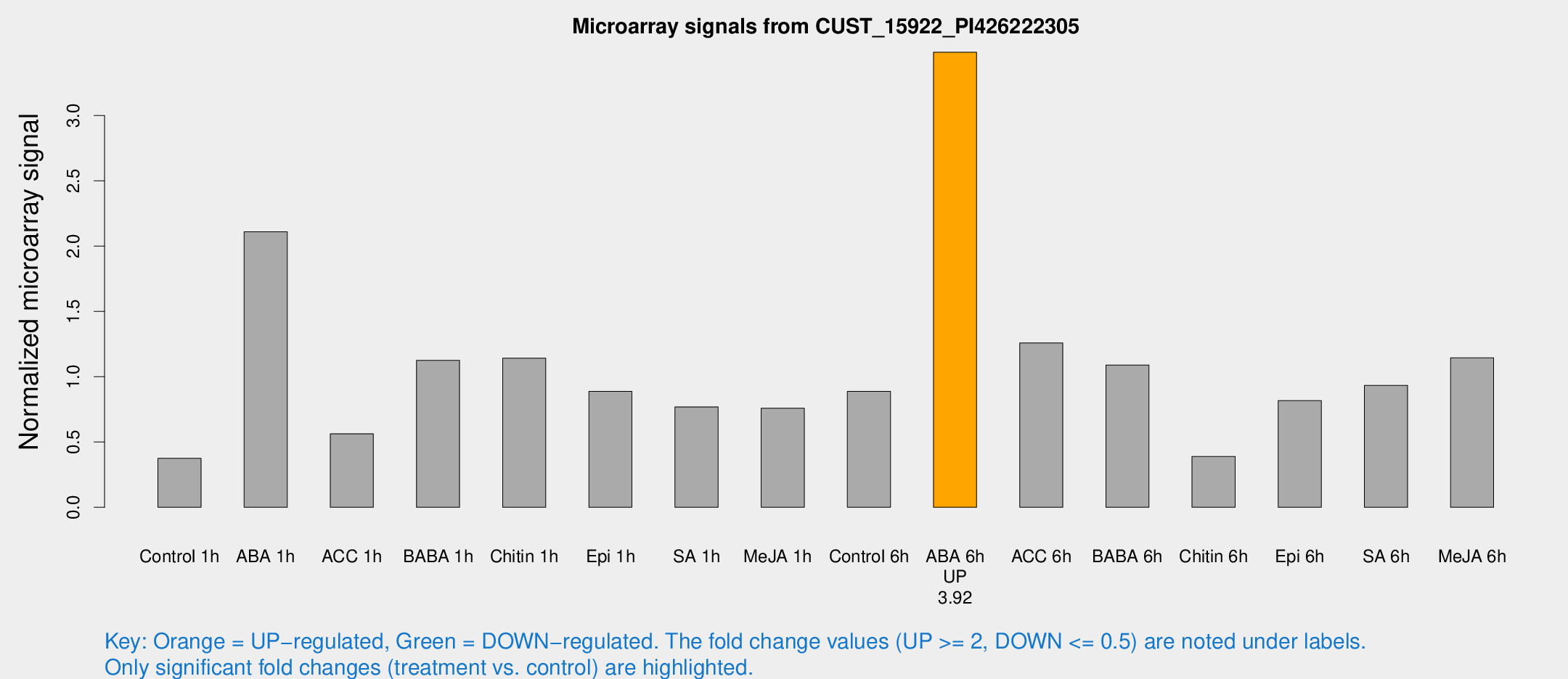
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.36894 | 3.20193 | 0.375341 | 0.195596 |
| ABA 1h | 31.9984 | 5.60879 | 2.11082 | 0.333654 |
| ACC 1h | 10.6397 | 3.94477 | 0.563097 | 0.255061 |
| BABA 1h | 18.8868 | 4.0239 | 1.1255 | 0.303328 |
| Chitin 1h | 17.6347 | 3.34073 | 1.14258 | 0.22389 |
| Epi 1h | 13.0031 | 3.19061 | 0.888566 | 0.233004 |
| SA 1h | 13.7888 | 3.31123 | 0.768828 | 0.210114 |
| Me-JA 1h | 12.0091 | 4.78911 | 0.75906 | 0.401812 |
| Control 6h | 16.392 | 4.78767 | 0.888328 | 0.250059 |
| ABA 6h | 63.1876 | 9.83836 | 3.48539 | 0.46142 |
| ACC 6h | 24.497 | 4.13303 | 1.25945 | 0.31993 |
| BABA 6h | 21.8569 | 5.65825 | 1.09001 | 0.281911 |
| Chitin 6h | 6.93251 | 3.62711 | 0.388915 | 0.207182 |
| Epi 6h | 15.3001 | 3.91306 | 0.816597 | 0.209346 |
| SA 6h | 20.574 | 10.4985 | 0.933234 | 0.700265 |
| Me-JA 6h | 19.1473 | 3.43148 | 1.14456 | 0.210848 |
Source Transcript PGSC0003DMT400076378 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G28260.1 | +3 | 4e-167 | 521 | 368/999 (37%) | Telomerase activating protein Est1 | chr1:9875776-9878782 REVERSE LENGTH=880 |
