Probe CUST_15892_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15892_PI426222305 | JHI_St_60k_v1 | DMT400076444 | GTTATTAATGCAAGTTATACTCTCGTGGGTGACACCATCTTCCATTCCATACATTCTAAG |
All Microarray Probes Designed to Gene DMG400029728
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15892_PI426222305 | JHI_St_60k_v1 | DMT400076444 | GTTATTAATGCAAGTTATACTCTCGTGGGTGACACCATCTTCCATTCCATACATTCTAAG |
Microarray Signals from CUST_15892_PI426222305
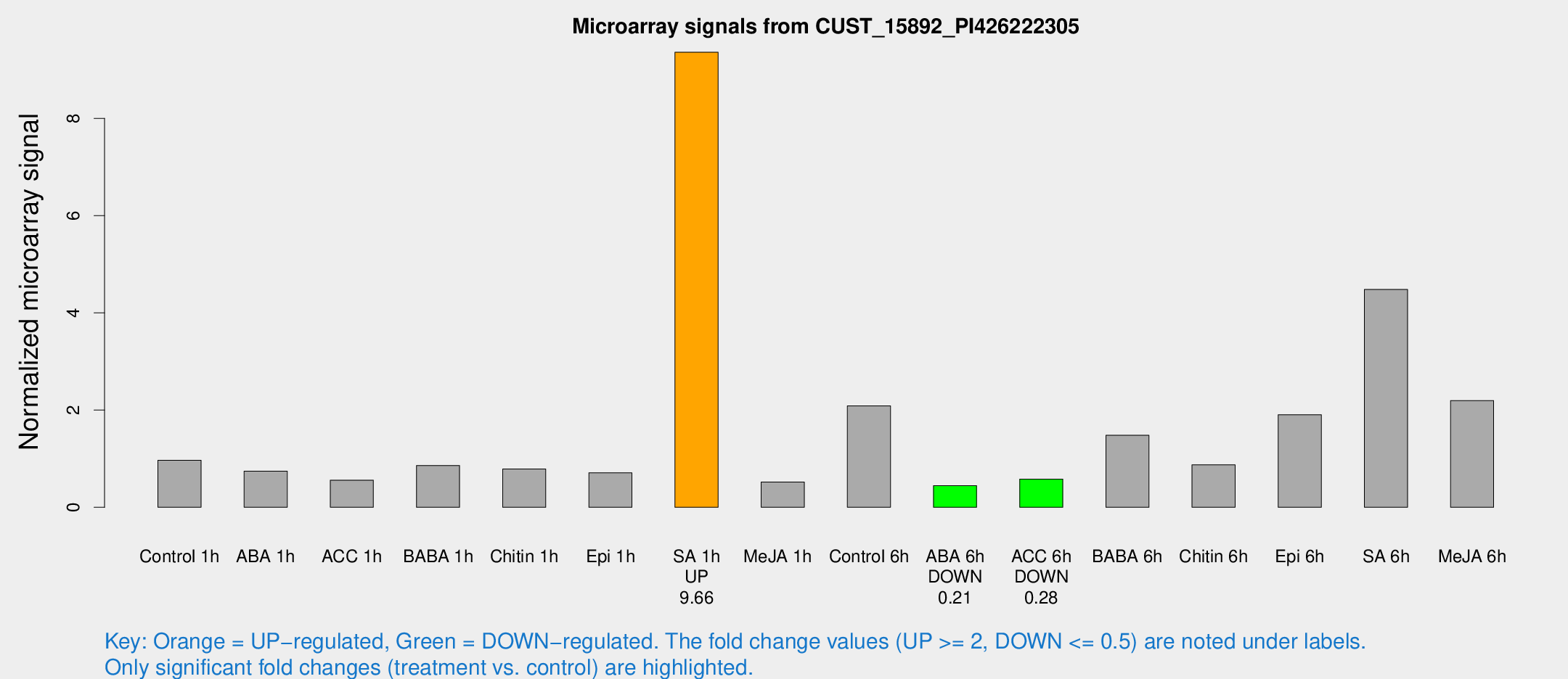
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 163.46 | 24.3014 | 0.968607 | 0.129437 |
| ABA 1h | 120.808 | 41.6582 | 0.742373 | 0.256746 |
| ACC 1h | 103.198 | 29.5285 | 0.558445 | 0.138769 |
| BABA 1h | 142.599 | 28.3858 | 0.85941 | 0.102641 |
| Chitin 1h | 118.854 | 15.2119 | 0.787728 | 0.0952262 |
| Epi 1h | 108.625 | 26.5091 | 0.709844 | 0.197582 |
| SA 1h | 1687.87 | 426.871 | 9.36084 | 1.77942 |
| Me-JA 1h | 72.1165 | 11.0703 | 0.521104 | 0.0547727 |
| Control 6h | 380.524 | 105.893 | 2.08569 | 0.532769 |
| ABA 6h | 78.3061 | 5.57517 | 0.446403 | 0.0317829 |
| ACC 6h | 110.769 | 8.96659 | 0.581166 | 0.0684338 |
| BABA 6h | 277.121 | 32.8797 | 1.48055 | 0.227175 |
| Chitin 6h | 163.527 | 42.6346 | 0.875162 | 0.196235 |
| Epi 6h | 374.027 | 82.6619 | 1.90479 | 0.313576 |
| SA 6h | 793.003 | 225.502 | 4.48088 | 0.89731 |
| Me-JA 6h | 403.677 | 128.036 | 2.19639 | 0.78015 |
Source Transcript PGSC0003DMT400076444 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G20970.1 | +1 | 4e-22 | 92 | 53/124 (43%) | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | chr4:11215259-11216212 FORWARD LENGTH=190 |
