Probe CUST_15824_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15824_PI426222305 | JHI_St_60k_v1 | DMT400057529 | GCAGATTTCACAGAAGAAGTGTAATTGGATTTTCACCTAACAGAACAGAGGAAAATATTC |
All Microarray Probes Designed to Gene DMG400022345
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15824_PI426222305 | JHI_St_60k_v1 | DMT400057529 | GCAGATTTCACAGAAGAAGTGTAATTGGATTTTCACCTAACAGAACAGAGGAAAATATTC |
Microarray Signals from CUST_15824_PI426222305
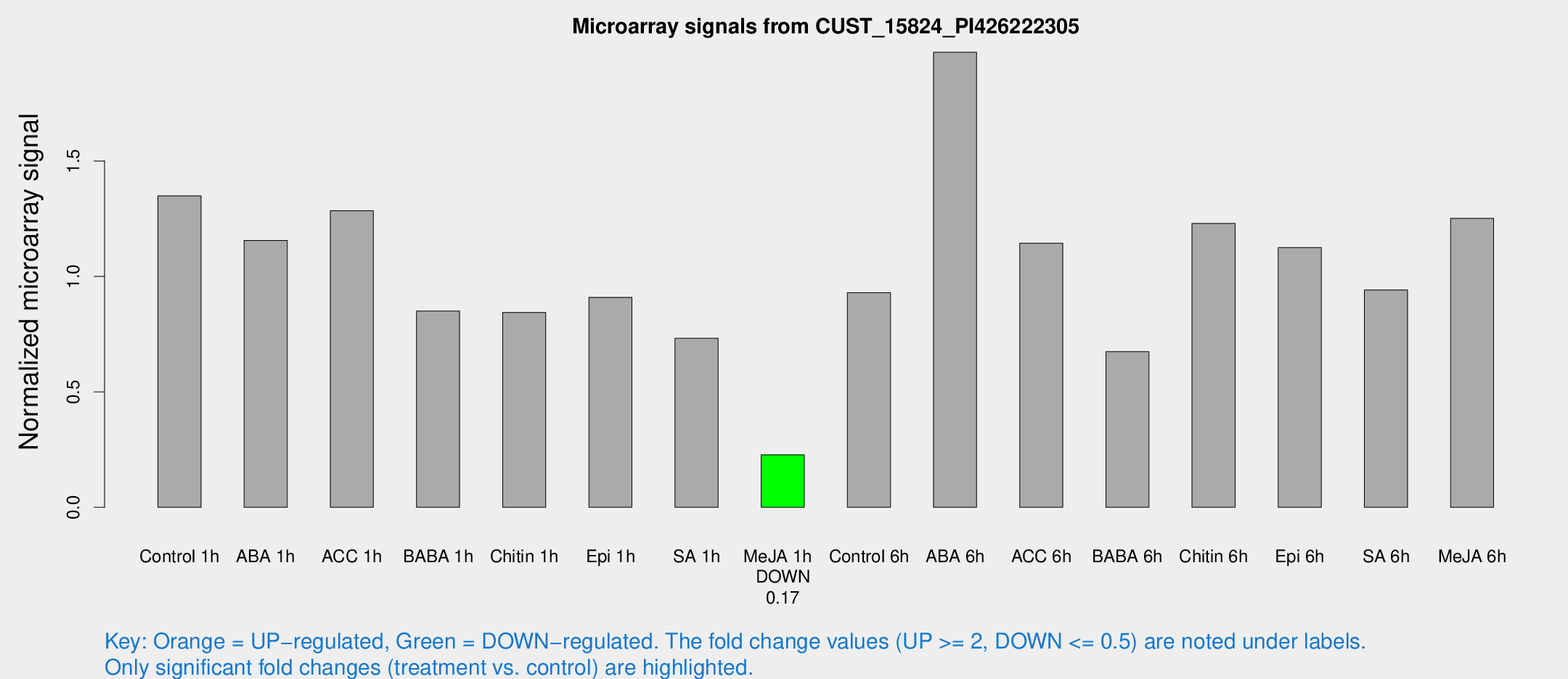
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1559.22 | 103.868 | 1.3488 | 0.077917 |
| ABA 1h | 1215.05 | 225.339 | 1.15619 | 0.221474 |
| ACC 1h | 1723.55 | 524.086 | 1.28489 | 0.423242 |
| BABA 1h | 994.901 | 211.376 | 0.849903 | 0.118565 |
| Chitin 1h | 933.407 | 252.526 | 0.844125 | 0.160265 |
| Epi 1h | 934.31 | 171.378 | 0.909291 | 0.169735 |
| SA 1h | 894.251 | 152.622 | 0.732183 | 0.108549 |
| Me-JA 1h | 215.726 | 19.1128 | 0.227661 | 0.0136335 |
| Control 6h | 1089.8 | 144.537 | 0.929222 | 0.110251 |
| ABA 6h | 2480.18 | 386.337 | 1.97123 | 0.251142 |
| ACC 6h | 1522.31 | 128.063 | 1.14431 | 0.263213 |
| BABA 6h | 895.1 | 151.897 | 0.674283 | 0.133858 |
| Chitin 6h | 1565.71 | 286.29 | 1.22934 | 0.300306 |
| Epi 6h | 1485.31 | 197.621 | 1.12552 | 0.0764544 |
| SA 6h | 1119.49 | 214.182 | 0.941693 | 0.0923794 |
| Me-JA 6h | 1657.86 | 621.207 | 1.2515 | 0.485125 |
Source Transcript PGSC0003DMT400057529 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G54470.1 | +1 | 7e-37 | 133 | 59/90 (66%) | B-box type zinc finger family protein | chr5:22114584-22115315 REVERSE LENGTH=215 |
