Probe CUST_15613_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15613_PI426222305 | JHI_St_60k_v1 | DMT400011285 | TCAGACCATTACCATCATCTCATCATCAAGGTTTTCAAGCAAATCTGGTTTGATATTCAA |
All Microarray Probes Designed to Gene DMG400004418
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15613_PI426222305 | JHI_St_60k_v1 | DMT400011285 | TCAGACCATTACCATCATCTCATCATCAAGGTTTTCAAGCAAATCTGGTTTGATATTCAA |
| CUST_15643_PI426222305 | JHI_St_60k_v1 | DMT400011286 | TCAAGTCGGAGCTAGAAGTTTGGGTATGGATTTGATCGAACCAGTAACTTTTGATCAAAT |
Microarray Signals from CUST_15613_PI426222305
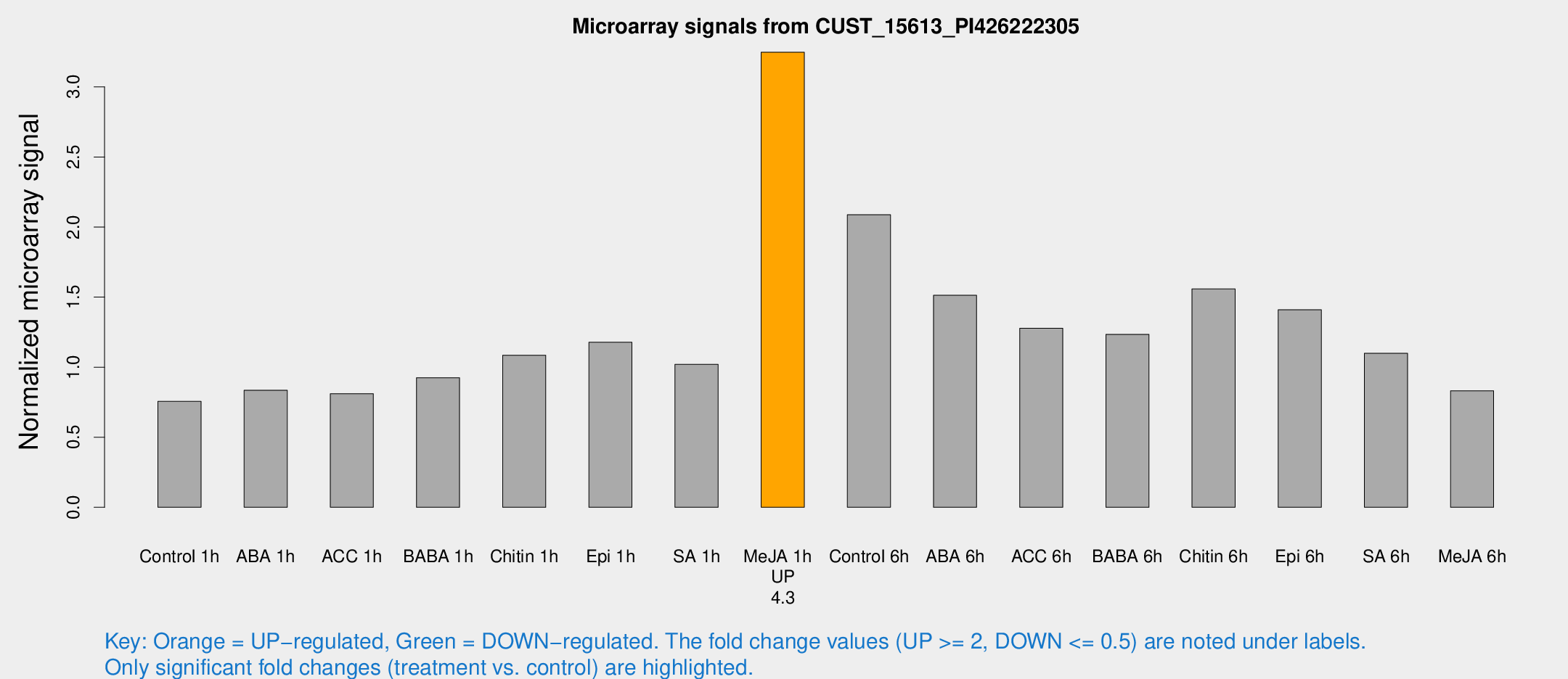
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.03884 | 3.46939 | 0.755757 | 0.434092 |
| ABA 1h | 5.90566 | 3.42453 | 0.835412 | 0.484039 |
| ACC 1h | 6.65342 | 3.86001 | 0.810634 | 0.469868 |
| BABA 1h | 7.16732 | 3.77607 | 0.924534 | 0.493187 |
| Chitin 1h | 7.85059 | 3.63495 | 1.08526 | 0.515602 |
| Epi 1h | 8.74972 | 3.51578 | 1.17787 | 0.559284 |
| SA 1h | 8.60641 | 3.53388 | 1.02042 | 0.449954 |
| Me-JA 1h | 21.1709 | 3.74777 | 3.24749 | 0.575379 |
| Control 6h | 19.8049 | 6.76229 | 2.08734 | 0.830838 |
| ABA 6h | 14.7691 | 5.35542 | 1.51304 | 0.703281 |
| ACC 6h | 13.4235 | 5.05515 | 1.27732 | 0.51651 |
| BABA 6h | 11.6914 | 4.31034 | 1.23419 | 0.511734 |
| Chitin 6h | 13.951 | 4.24819 | 1.55841 | 0.536274 |
| Epi 6h | 13.0927 | 4.37711 | 1.40964 | 0.532528 |
| SA 6h | 9.21785 | 4.01019 | 1.09907 | 0.537628 |
| Me-JA 6h | 6.62112 | 3.84354 | 0.831814 | 0.482333 |
Source Transcript PGSC0003DMT400011285 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G01840.1 | +1 | 2e-25 | 100 | 52/89 (58%) | ovate family protein 1 | chr5:324552-325364 FORWARD LENGTH=270 |
