Probe CUST_15479_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15479_PI426222305 | JHI_St_60k_v1 | DMT400037473 | CTATCATCTTAGAGGGATCTGTTCTTAATTCAAATGGACAACAAGTCTTATGTGATGCAT |
All Microarray Probes Designed to Gene DMG400014459
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15479_PI426222305 | JHI_St_60k_v1 | DMT400037473 | CTATCATCTTAGAGGGATCTGTTCTTAATTCAAATGGACAACAAGTCTTATGTGATGCAT |
Microarray Signals from CUST_15479_PI426222305
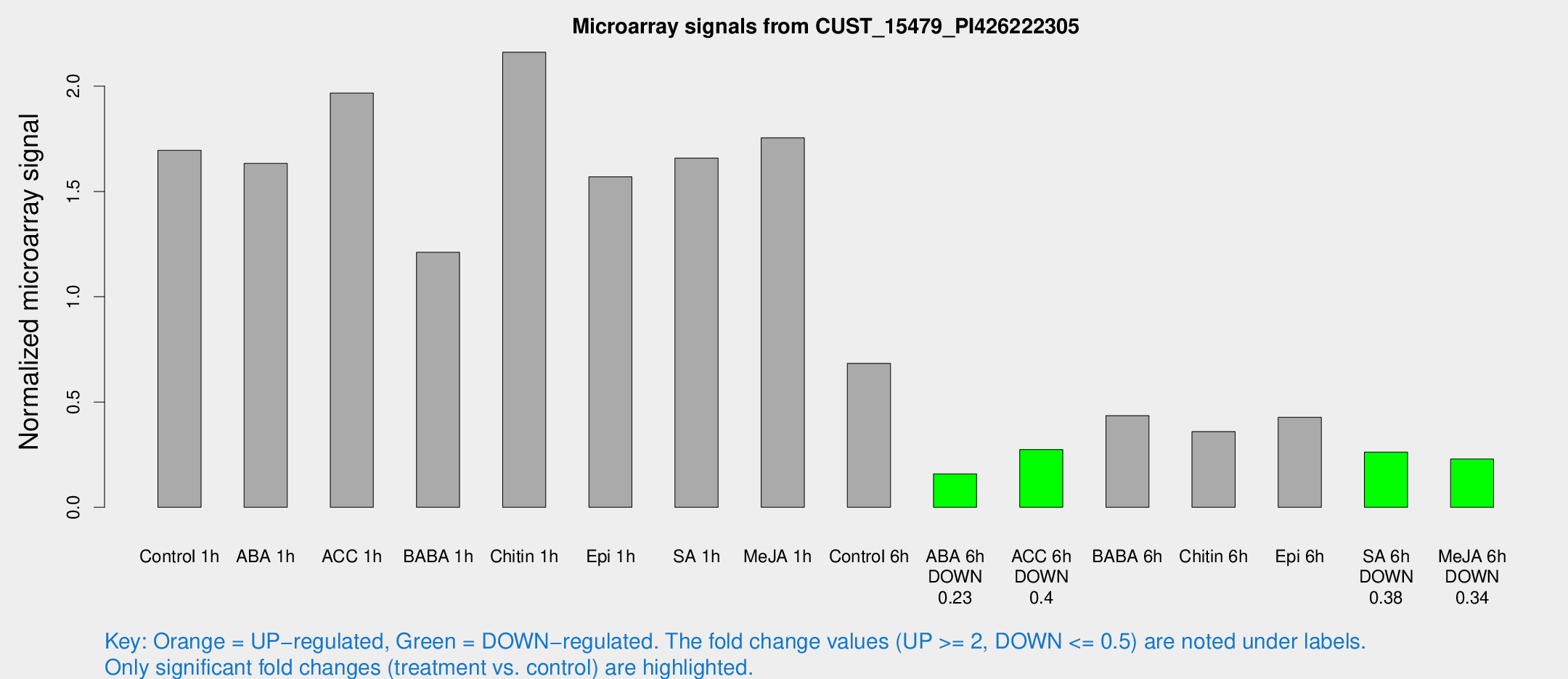
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11563 | 1185.79 | 1.69502 | 0.120483 |
| ABA 1h | 9745.42 | 562.764 | 1.63307 | 0.109859 |
| ACC 1h | 14207.7 | 3038.01 | 1.96735 | 0.581853 |
| BABA 1h | 8142.71 | 1452.86 | 1.21094 | 0.118333 |
| Chitin 1h | 13494 | 2370.63 | 2.16101 | 0.368413 |
| Epi 1h | 9170.7 | 529.887 | 1.56968 | 0.0906274 |
| SA 1h | 11656.9 | 1455.99 | 1.65796 | 0.125095 |
| Me-JA 1h | 9938.35 | 1669.88 | 1.75413 | 0.264766 |
| Control 6h | 5024.42 | 1324.76 | 0.683366 | 0.146573 |
| ABA 6h | 1139.03 | 66.0097 | 0.158513 | 0.0091672 |
| ACC 6h | 2265.07 | 535.363 | 0.274433 | 0.0470289 |
| BABA 6h | 3357.94 | 467.479 | 0.43518 | 0.0830699 |
| Chitin 6h | 2586.13 | 149.658 | 0.359498 | 0.0207631 |
| Epi 6h | 3253.67 | 188.03 | 0.427525 | 0.0405153 |
| SA 6h | 1942.03 | 551.51 | 0.26219 | 0.0621761 |
| Me-JA 6h | 1617.34 | 378.029 | 0.229058 | 0.0348539 |
Source Transcript PGSC0003DMT400037473 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G48600.2 | +1 | 0.0 | 797 | 385/485 (79%) | S-adenosyl-L-methionine-dependent methyltransferases superfamily protein | chr1:17966074-17969077 FORWARD LENGTH=491 |
