Probe CUST_14689_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14689_PI426222305 | JHI_St_60k_v1 | DMT400066595 | TTGCTCCAAGTACTTTGACACATGGTATCTAATAAGCCAACACGAGATGATAAGTTTAAC |
All Microarray Probes Designed to Gene DMG400025887
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14689_PI426222305 | JHI_St_60k_v1 | DMT400066595 | TTGCTCCAAGTACTTTGACACATGGTATCTAATAAGCCAACACGAGATGATAAGTTTAAC |
| CUST_14604_PI426222305 | JHI_St_60k_v1 | DMT400066596 | TTGCTCCAAGTACTTTGACACATGGTATCTAATAAGCCAACACGAGATGATAAGTTTAAC |
Microarray Signals from CUST_14689_PI426222305
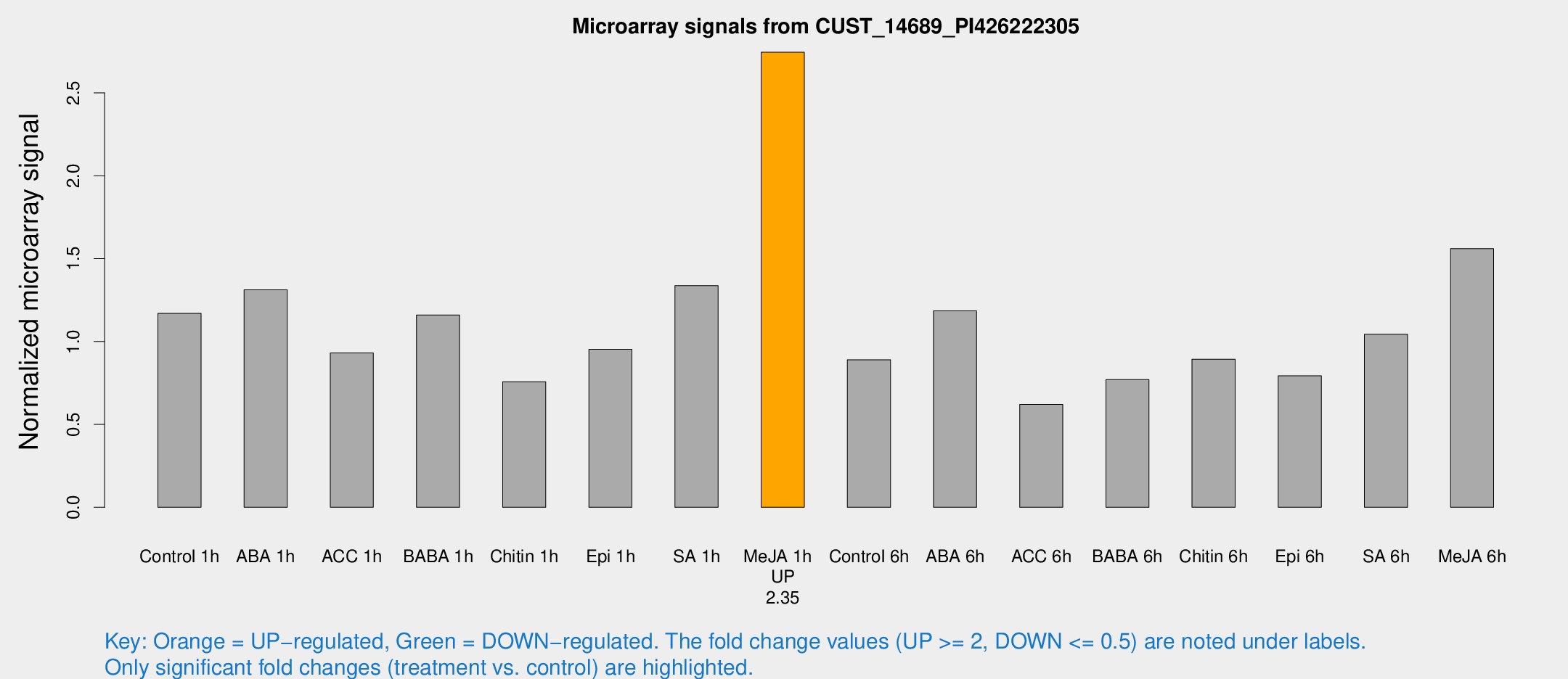
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 122.254 | 13.0771 | 1.17048 | 0.101242 |
| ABA 1h | 132.04 | 39.8245 | 1.31204 | 0.338397 |
| ACC 1h | 105.431 | 28.3961 | 0.930976 | 0.219078 |
| BABA 1h | 122.14 | 25.9247 | 1.16034 | 0.166694 |
| Chitin 1h | 71.1387 | 7.5761 | 0.757888 | 0.0564335 |
| Epi 1h | 85.6242 | 5.93324 | 0.953619 | 0.0663037 |
| SA 1h | 143.178 | 13.8288 | 1.33692 | 0.10955 |
| Me-JA 1h | 233.331 | 20.9521 | 2.74559 | 0.16373 |
| Control 6h | 95.5477 | 18.3096 | 0.889552 | 0.099112 |
| ABA 6h | 130.928 | 10.0388 | 1.18565 | 0.0757748 |
| ACC 6h | 73.9547 | 6.01829 | 0.620558 | 0.0705258 |
| BABA 6h | 89.2073 | 6.41462 | 0.770918 | 0.055505 |
| Chitin 6h | 98.3188 | 6.8244 | 0.893726 | 0.0620624 |
| Epi 6h | 93.0188 | 7.88874 | 0.793232 | 0.129475 |
| SA 6h | 113.041 | 24.4543 | 1.04403 | 0.14161 |
| Me-JA 6h | 163.367 | 22.5286 | 1.56023 | 0.102069 |
Source Transcript PGSC0003DMT400066595 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G15570.2 | +1 | 3e-50 | 165 | 74/106 (70%) | Thioredoxin superfamily protein | chr2:6791556-6792902 REVERSE LENGTH=174 |
