Probe CUST_14445_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14445_PI426222305 | JHI_St_60k_v1 | DMT400066568 | GTTAAGACAATTTGGACTTTCTTTGCTATATAATGCTAGCGTTTGGCCATAGATTGACAA |
All Microarray Probes Designed to Gene DMG400025872
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14445_PI426222305 | JHI_St_60k_v1 | DMT400066568 | GTTAAGACAATTTGGACTTTCTTTGCTATATAATGCTAGCGTTTGGCCATAGATTGACAA |
Microarray Signals from CUST_14445_PI426222305
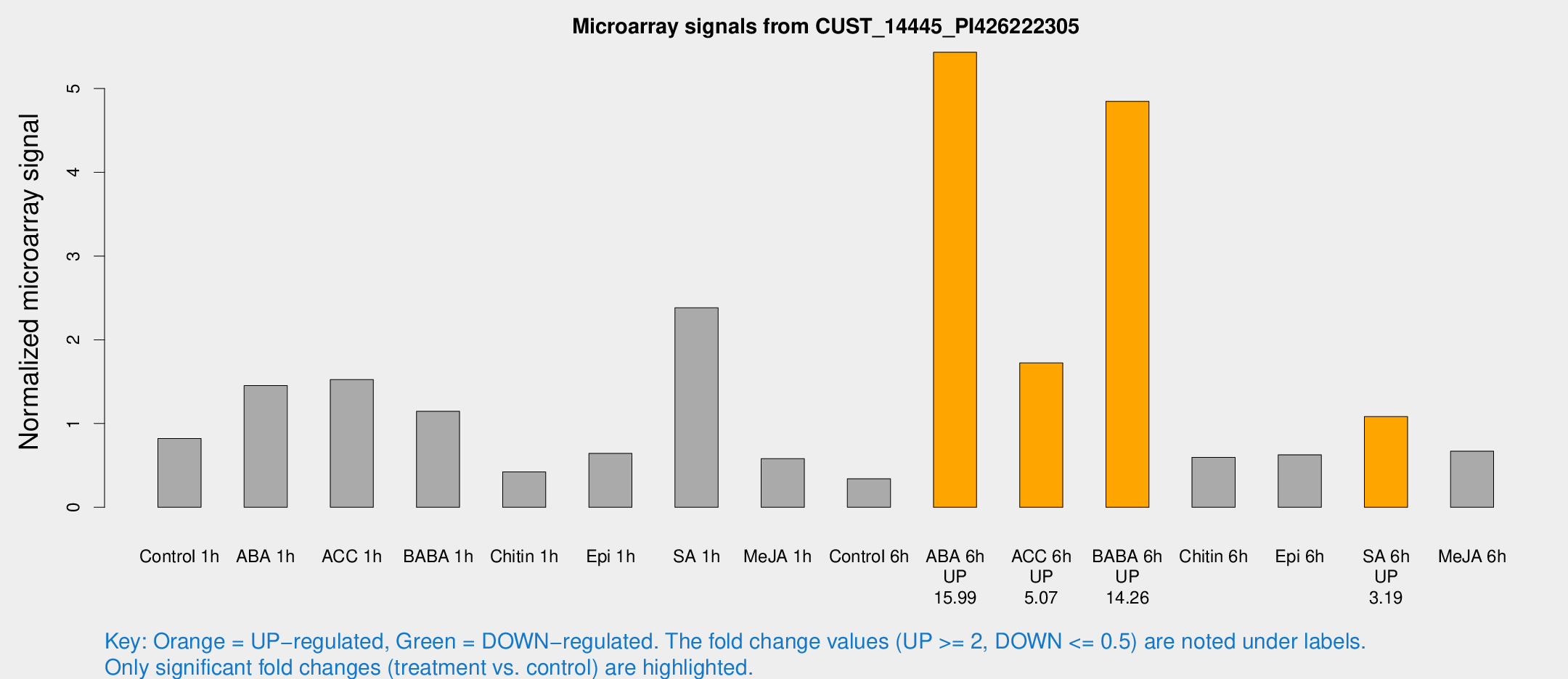
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 107.606 | 40.6965 | 0.821222 | 0.256011 |
| ABA 1h | 200.405 | 93.7653 | 1.45379 | 1.06378 |
| ACC 1h | 246.142 | 103.727 | 1.52491 | 1.11843 |
| BABA 1h | 144.294 | 43.3761 | 1.1465 | 0.346842 |
| Chitin 1h | 51.7792 | 22.4299 | 0.423408 | 0.152363 |
| Epi 1h | 68.308 | 17.4149 | 0.643321 | 0.175125 |
| SA 1h | 339.875 | 152.252 | 2.3819 | 0.975974 |
| Me-JA 1h | 63.7131 | 23.737 | 0.580634 | 0.205164 |
| Control 6h | 40.8567 | 8.23624 | 0.339793 | 0.0425711 |
| ABA 6h | 735.633 | 225.556 | 5.43305 | 1.60123 |
| ACC 6h | 227.685 | 14.5604 | 1.72376 | 0.130202 |
| BABA 6h | 670.704 | 184.12 | 4.84641 | 1.32938 |
| Chitin 6h | 75.5908 | 14.8331 | 0.595502 | 0.111518 |
| Epi 6h | 88.9631 | 26.7048 | 0.626691 | 0.151526 |
| SA 6h | 125.473 | 17.6068 | 1.08249 | 0.0725358 |
| Me-JA 6h | 83.2501 | 25.0483 | 0.670405 | 0.143396 |
Source Transcript PGSC0003DMT400066568 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G34131.1 | +3 | 1e-154 | 454 | 230/481 (48%) | UDP-glucosyl transferase 73B3 | chr4:16343268-16344713 REVERSE LENGTH=481 |
