Probe CUST_14166_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14166_PI426222305 | JHI_St_60k_v1 | DMT400060057 | GATTAAAAACCTAGATATATACTTGATGTCAAAGTACTAAGGGGGATTACCCTCATGATG |
All Microarray Probes Designed to Gene DMG400023365
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14166_PI426222305 | JHI_St_60k_v1 | DMT400060057 | GATTAAAAACCTAGATATATACTTGATGTCAAAGTACTAAGGGGGATTACCCTCATGATG |
Microarray Signals from CUST_14166_PI426222305
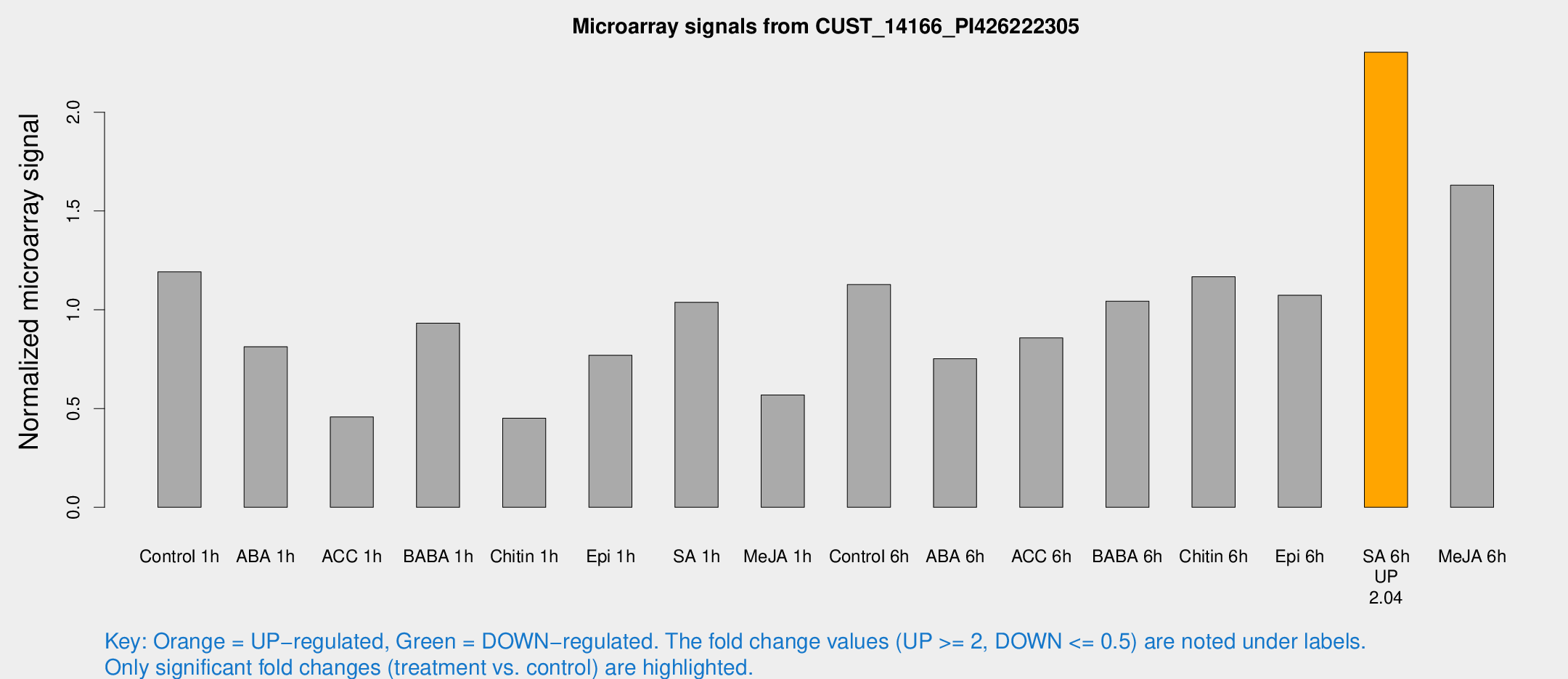
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 933.919 | 226.627 | 1.19224 | 0.205874 |
| ABA 1h | 580.964 | 154.198 | 0.812347 | 0.211378 |
| ACC 1h | 568.29 | 253.993 | 0.457708 | 0.600297 |
| BABA 1h | 765.01 | 236.093 | 0.931766 | 0.301731 |
| Chitin 1h | 312.797 | 62.5338 | 0.451051 | 0.0818059 |
| Epi 1h | 500.807 | 61.19 | 0.76997 | 0.0756425 |
| SA 1h | 838.369 | 204.437 | 1.03784 | 0.195843 |
| Me-JA 1h | 384.436 | 130.442 | 0.568975 | 0.170166 |
| Control 6h | 839.562 | 48.8103 | 1.1275 | 0.147878 |
| ABA 6h | 594.147 | 36.6202 | 0.752029 | 0.0436422 |
| ACC 6h | 736.704 | 77.277 | 0.857676 | 0.133995 |
| BABA 6h | 907.766 | 194.163 | 1.04361 | 0.210386 |
| Chitin 6h | 921.089 | 53.3761 | 1.1671 | 0.0791599 |
| Epi 6h | 899.958 | 59.8455 | 1.07329 | 0.129861 |
| SA 6h | 1695.55 | 116.993 | 2.30365 | 0.133091 |
| Me-JA 6h | 1300.78 | 319.619 | 1.63047 | 0.375494 |
Source Transcript PGSC0003DMT400060057 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G65480.1 | +1 | 2e-92 | 274 | 125/168 (74%) | PEBP (phosphatidylethanolamine-binding protein) family protein | chr1:24331510-24333689 FORWARD LENGTH=175 |
