Probe CUST_13148_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13148_PI426222305 | JHI_St_60k_v1 | DMT400007495 | GTGATGAGCCATTCTGATCTATTCCTGTTTTTGCTTATAGGGCTCGTGAAGATACTCAAG |
All Microarray Probes Designed to Gene DMG400002890
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13191_PI426222305 | JHI_St_60k_v1 | DMT400007496 | ATGGTTTTGCTGGAAAATTACAGTGCCCTCAACTATACTGGGCTCGTGAAGATACTCAAG |
| CUST_13148_PI426222305 | JHI_St_60k_v1 | DMT400007495 | GTGATGAGCCATTCTGATCTATTCCTGTTTTTGCTTATAGGGCTCGTGAAGATACTCAAG |
Microarray Signals from CUST_13148_PI426222305
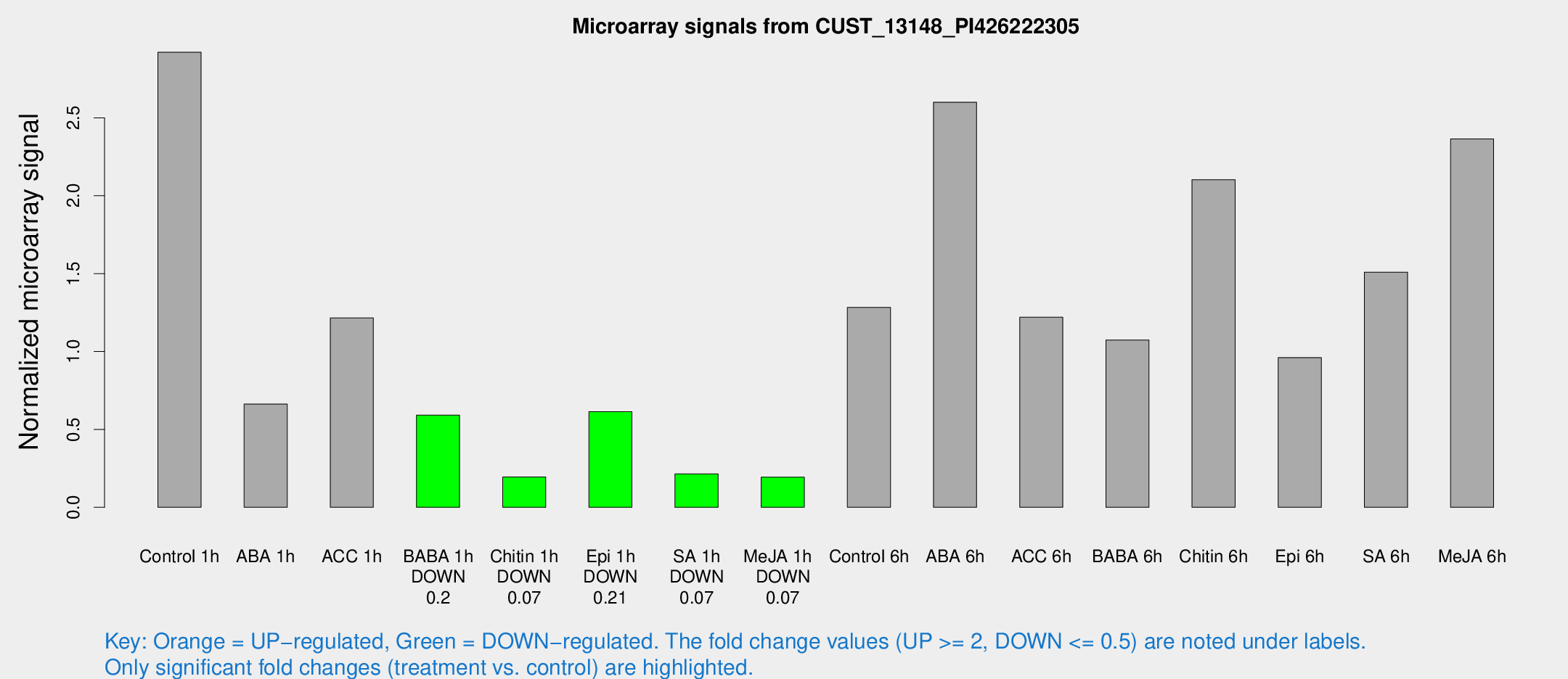
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 103.496 | 21.3785 | 2.92093 | 0.547961 |
| ABA 1h | 29.8441 | 15.6646 | 0.662332 | 0.669082 |
| ACC 1h | 54.9757 | 22.6816 | 1.21543 | 0.743235 |
| BABA 1h | 22.3439 | 7.98846 | 0.5913 | 0.200708 |
| Chitin 1h | 5.94229 | 2.85948 | 0.194292 | 0.0970949 |
| Epi 1h | 18.9495 | 4.80686 | 0.614115 | 0.132482 |
| SA 1h | 8.36384 | 3.14759 | 0.2144 | 0.0948907 |
| Me-JA 1h | 5.1432 | 2.91197 | 0.193181 | 0.105433 |
| Control 6h | 64.604 | 41.1353 | 1.28306 | 0.988927 |
| ABA 6h | 112.136 | 38.8203 | 2.6001 | 1.29537 |
| ACC 6h | 54.6084 | 21.421 | 1.21998 | 0.313996 |
| BABA 6h | 52.7947 | 25.8442 | 1.07324 | 0.665592 |
| Chitin 6h | 77.7182 | 13.4176 | 2.10247 | 0.421462 |
| Epi 6h | 40.5159 | 11.256 | 0.960756 | 0.285213 |
| SA 6h | 56.9529 | 19.5231 | 1.50906 | 0.641139 |
| Me-JA 6h | 88.6482 | 28.1195 | 2.3647 | 0.64769 |
Source Transcript PGSC0003DMT400007495 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G26660.1 | +1 | 2e-48 | 172 | 98/151 (65%) | SPX domain gene 2 | chr2:11338932-11340703 FORWARD LENGTH=287 |
