Probe CUST_13102_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13102_PI426222305 | JHI_St_60k_v1 | DMT400007507 | AGTCGATGGATGAACTTGTCGAGTTTTATTCATGTTGATTGATTCAGAATGAACGTCATC |
All Microarray Probes Designed to Gene DMG400002896
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13102_PI426222305 | JHI_St_60k_v1 | DMT400007507 | AGTCGATGGATGAACTTGTCGAGTTTTATTCATGTTGATTGATTCAGAATGAACGTCATC |
Microarray Signals from CUST_13102_PI426222305
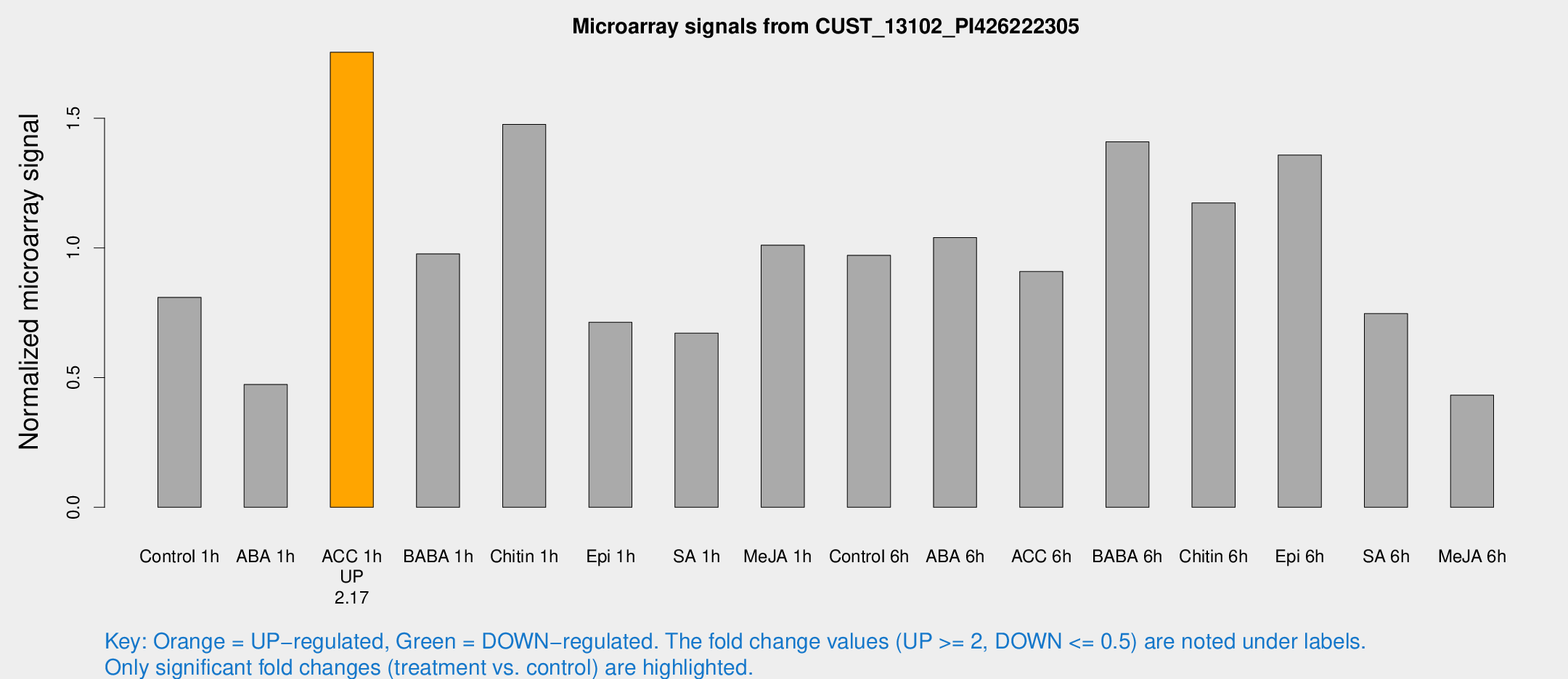
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 15.99 | 3.7163 | 0.809392 | 0.191077 |
| ABA 1h | 9.08591 | 3.59357 | 0.473316 | 0.225916 |
| ACC 1h | 35.62 | 4.58027 | 1.75426 | 0.231519 |
| BABA 1h | 19.9885 | 5.07671 | 0.976908 | 0.367069 |
| Chitin 1h | 34.8235 | 18.8571 | 1.476 | 1.06028 |
| Epi 1h | 13.839 | 4.64815 | 0.713381 | 0.289358 |
| SA 1h | 15.3074 | 4.56248 | 0.671165 | 0.336001 |
| Me-JA 1h | 16.3935 | 3.87021 | 1.01049 | 0.248922 |
| Control 6h | 23.7829 | 8.58497 | 0.971453 | 0.485338 |
| ABA 6h | 31.5347 | 17.8308 | 1.03996 | 0.9982 |
| ACC 6h | 22.6327 | 6.35215 | 0.908778 | 0.23947 |
| BABA 6h | 33.8362 | 10.2884 | 1.40894 | 0.386122 |
| Chitin 6h | 25.5439 | 5.45162 | 1.17315 | 0.259596 |
| Epi 6h | 32.2107 | 8.92889 | 1.35778 | 0.398757 |
| SA 6h | 16.824 | 6.65346 | 0.746743 | 0.254599 |
| Me-JA 6h | 8.62738 | 3.76889 | 0.432452 | 0.19776 |
Source Transcript PGSC0003DMT400007507 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G20820.1 | +2 | 3e-30 | 110 | 64/134 (48%) | SAUR-like auxin-responsive protein family | chr5:7046911-7047294 REVERSE LENGTH=127 |
