Probe CUST_13080_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13080_PI426222305 | JHI_St_60k_v1 | DMT400007514 | ACCGCAAGAGAGTCAAAAGTTCGTGTCAAGTCAAAACTAGACAAACAAATTCAAACGAAG |
All Microarray Probes Designed to Gene DMG400002901
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13188_PI426222305 | JHI_St_60k_v1 | DMT400007516 | TAACACCGCAAGAGAGTCAAAAGTTCGTGTCAAGTCAAAACTAGACAAACAAATTCAAAC |
| CUST_13080_PI426222305 | JHI_St_60k_v1 | DMT400007514 | ACCGCAAGAGAGTCAAAAGTTCGTGTCAAGTCAAAACTAGACAAACAAATTCAAACGAAG |
Microarray Signals from CUST_13080_PI426222305
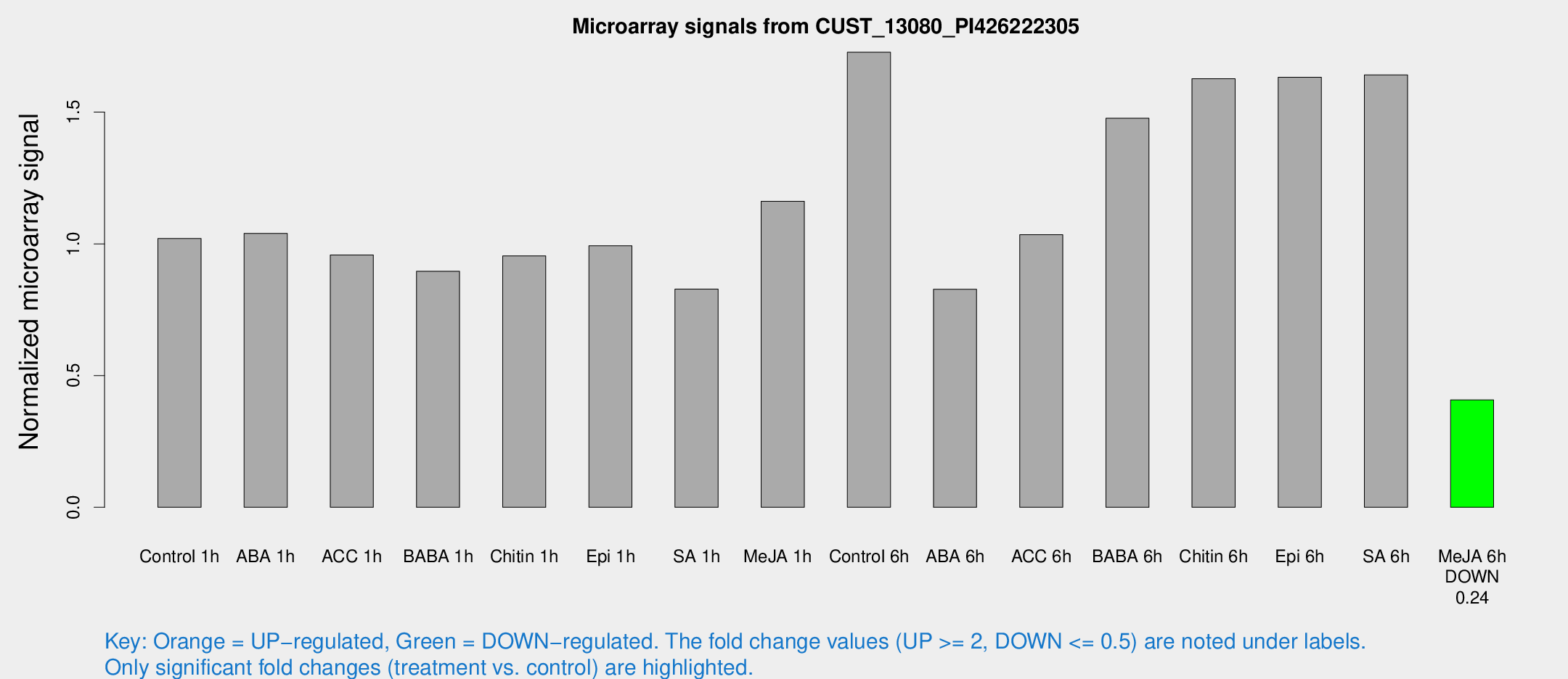
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 39.9378 | 5.321 | 1.02017 | 0.118945 |
| ABA 1h | 36.283 | 5.60031 | 1.03953 | 0.137567 |
| ACC 1h | 38.3596 | 4.46465 | 0.957612 | 0.118723 |
| BABA 1h | 33.7663 | 4.39331 | 0.896057 | 0.113228 |
| Chitin 1h | 33.397 | 3.94571 | 0.954307 | 0.124422 |
| Epi 1h | 33.4557 | 3.96275 | 0.992561 | 0.119108 |
| SA 1h | 32.9763 | 3.94497 | 0.82844 | 0.100316 |
| Me-JA 1h | 36.8277 | 4.28405 | 1.16115 | 0.14628 |
| Control 6h | 71.6015 | 17.6338 | 1.72724 | 0.350377 |
| ABA 6h | 33.9929 | 4.16228 | 0.827939 | 0.102247 |
| ACC 6h | 47.3755 | 8.43221 | 1.03479 | 0.173225 |
| BABA 6h | 67.243 | 15.14 | 1.47654 | 0.414498 |
| Chitin 6h | 67.1804 | 6.19457 | 1.62667 | 0.136601 |
| Epi 6h | 70.8546 | 5.85255 | 1.63221 | 0.134184 |
| SA 6h | 63.9645 | 9.49641 | 1.64137 | 0.141534 |
| Me-JA 6h | 19.5407 | 9.53499 | 0.407586 | 0.176128 |
Source Transcript PGSC0003DMT400007514 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G19150.1 | +1 | 7e-145 | 415 | 190/220 (86%) | photosystem I light harvesting complex gene 6 | chr1:6612806-6613799 FORWARD LENGTH=270 |
