Probe CUST_12956_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12956_PI426222305 | JHI_St_60k_v1 | DMT400062960 | GGTCTTGCTTTGTTTTCATATTCATTGATATATTTATTGTGCCCAGTTGTTGCAACTCTG |
All Microarray Probes Designed to Gene DMG400024501
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12980_PI426222305 | JHI_St_60k_v1 | DMT400062961 | CTGTTTTCCAACTACATGGTGATCTTGATACGCTTTGCAGGTTGGATGAGGCTATTGAAA |
| CUST_12956_PI426222305 | JHI_St_60k_v1 | DMT400062960 | GGTCTTGCTTTGTTTTCATATTCATTGATATATTTATTGTGCCCAGTTGTTGCAACTCTG |
Microarray Signals from CUST_12956_PI426222305
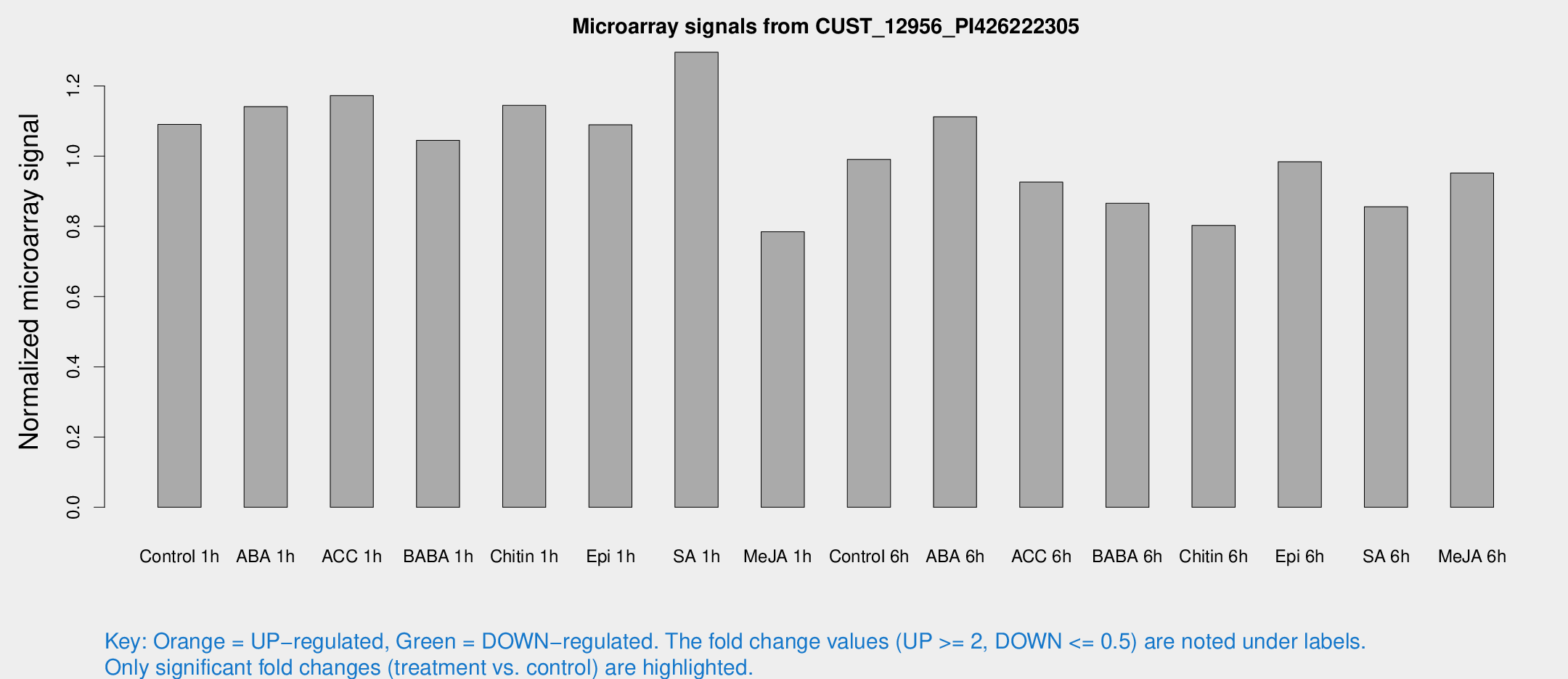
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2769.4 | 552.521 | 1.09019 | 0.140043 |
| ABA 1h | 2485.46 | 198.429 | 1.14106 | 0.102851 |
| ACC 1h | 3024.43 | 467.563 | 1.17255 | 0.103388 |
| BABA 1h | 2574.35 | 496.721 | 1.04487 | 0.119258 |
| Chitin 1h | 2544.01 | 263.334 | 1.14461 | 0.121674 |
| Epi 1h | 2339.75 | 292.258 | 1.08906 | 0.102098 |
| SA 1h | 3305.61 | 402.589 | 1.29582 | 0.0790603 |
| Me-JA 1h | 1596.29 | 210.517 | 0.784626 | 0.0453426 |
| Control 6h | 2665.02 | 711.427 | 0.990821 | 0.233706 |
| ABA 6h | 2958.36 | 461.748 | 1.11204 | 0.11948 |
| ACC 6h | 2648.34 | 358.696 | 0.92609 | 0.0571381 |
| BABA 6h | 2387.73 | 196.774 | 0.86573 | 0.0500057 |
| Chitin 6h | 2094.21 | 121.222 | 0.802719 | 0.0463734 |
| Epi 6h | 2741.65 | 270.966 | 0.984138 | 0.154108 |
| SA 6h | 2208.63 | 505.315 | 0.855735 | 0.130398 |
| Me-JA 6h | 2400.33 | 413.469 | 0.95173 | 0.117654 |
Source Transcript PGSC0003DMT400062960 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G27500.1 | +2 | 0.0 | 771 | 417/666 (63%) | Tetratricopeptide repeat (TPR)-like superfamily protein | chr1:9551629-9553654 REVERSE LENGTH=650 |
