Probe CUST_12441_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12441_PI426222305 | JHI_St_60k_v1 | DMT400007125 | TTACTCAAGGGGGAGAAAGAGATTGTTAGAAACTTCCTACTTCACACTCTCCAATTGCAG |
All Microarray Probes Designed to Gene DMG400002756
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12441_PI426222305 | JHI_St_60k_v1 | DMT400007125 | TTACTCAAGGGGGAGAAAGAGATTGTTAGAAACTTCCTACTTCACACTCTCCAATTGCAG |
| CUST_12516_PI426222305 | JHI_St_60k_v1 | DMT400007126 | TACTGTAACAAACATCGCTAGCTTGCTCCTACCTTTGCTATTTTTTGATGTTGCCCTTTC |
Microarray Signals from CUST_12441_PI426222305
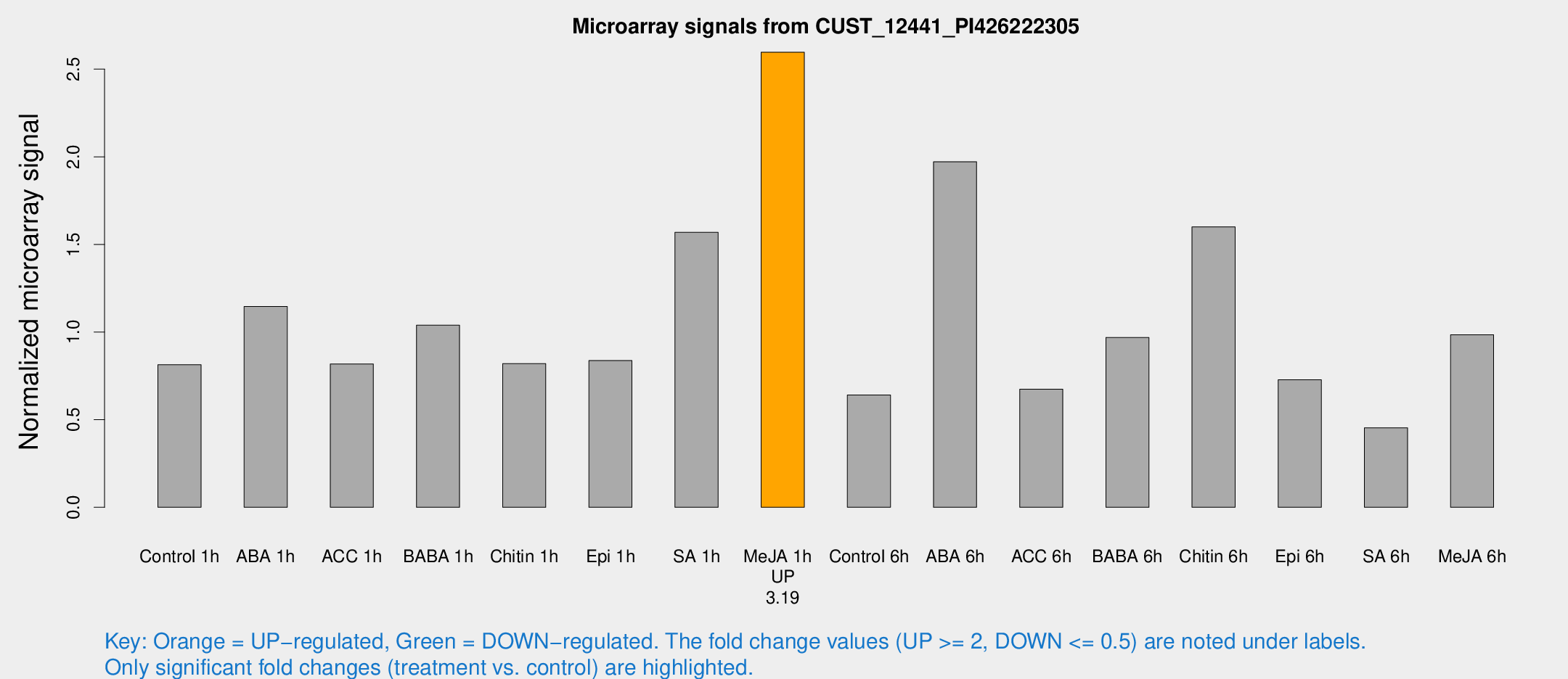
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 54.6249 | 10.4143 | 0.813594 | 0.110075 |
| ABA 1h | 67.7777 | 12.0882 | 1.14622 | 0.127502 |
| ACC 1h | 57.313 | 12.0349 | 0.8176 | 0.134102 |
| BABA 1h | 71.2153 | 18.6337 | 1.0392 | 0.232196 |
| Chitin 1h | 48.0017 | 4.36388 | 0.819716 | 0.0947557 |
| Epi 1h | 53.7448 | 18.0173 | 0.837183 | 0.331531 |
| SA 1h | 104.955 | 6.97645 | 1.56894 | 0.104195 |
| Me-JA 1h | 140.341 | 20.1909 | 2.59666 | 0.164252 |
| Control 6h | 53.9431 | 22.4915 | 0.640935 | 0.360955 |
| ABA 6h | 163.472 | 63.7871 | 1.97155 | 0.878612 |
| ACC 6h | 53.3829 | 12.139 | 0.673627 | 0.164083 |
| BABA 6h | 73.9922 | 15.6853 | 0.968934 | 0.186723 |
| Chitin 6h | 204.241 | 152.048 | 1.60036 | 2.02894 |
| Epi 6h | 58.8042 | 19.1462 | 0.728042 | 0.256241 |
| SA 6h | 29.1644 | 4.08074 | 0.453244 | 0.0639661 |
| Me-JA 6h | 75.2883 | 24.825 | 0.984597 | 0.414037 |
Source Transcript PGSC0003DMT400007125 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G56560.1 | +2 | 0.0 | 892 | 434/547 (79%) | Plant neutral invertase family protein | chr1:21192593-21194948 FORWARD LENGTH=616 |
