Probe CUST_11978_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11978_PI426222305 | JHI_St_60k_v1 | DMT400005956 | GTGCTTTGGCACAGTAGGGAAAGGGTGTTTACCAATTCGTGAAGAAATATGGAGCTAATG |
All Microarray Probes Designed to Gene DMG400002312
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11978_PI426222305 | JHI_St_60k_v1 | DMT400005956 | GTGCTTTGGCACAGTAGGGAAAGGGTGTTTACCAATTCGTGAAGAAATATGGAGCTAATG |
| CUST_12052_PI426222305 | JHI_St_60k_v1 | DMT400005957 | TCTGGCAGGAAGGCCAAGCGTCAAGCAAAGTTTTAGGAAAAATAATAACTTTTTAGCAAA |
Microarray Signals from CUST_11978_PI426222305
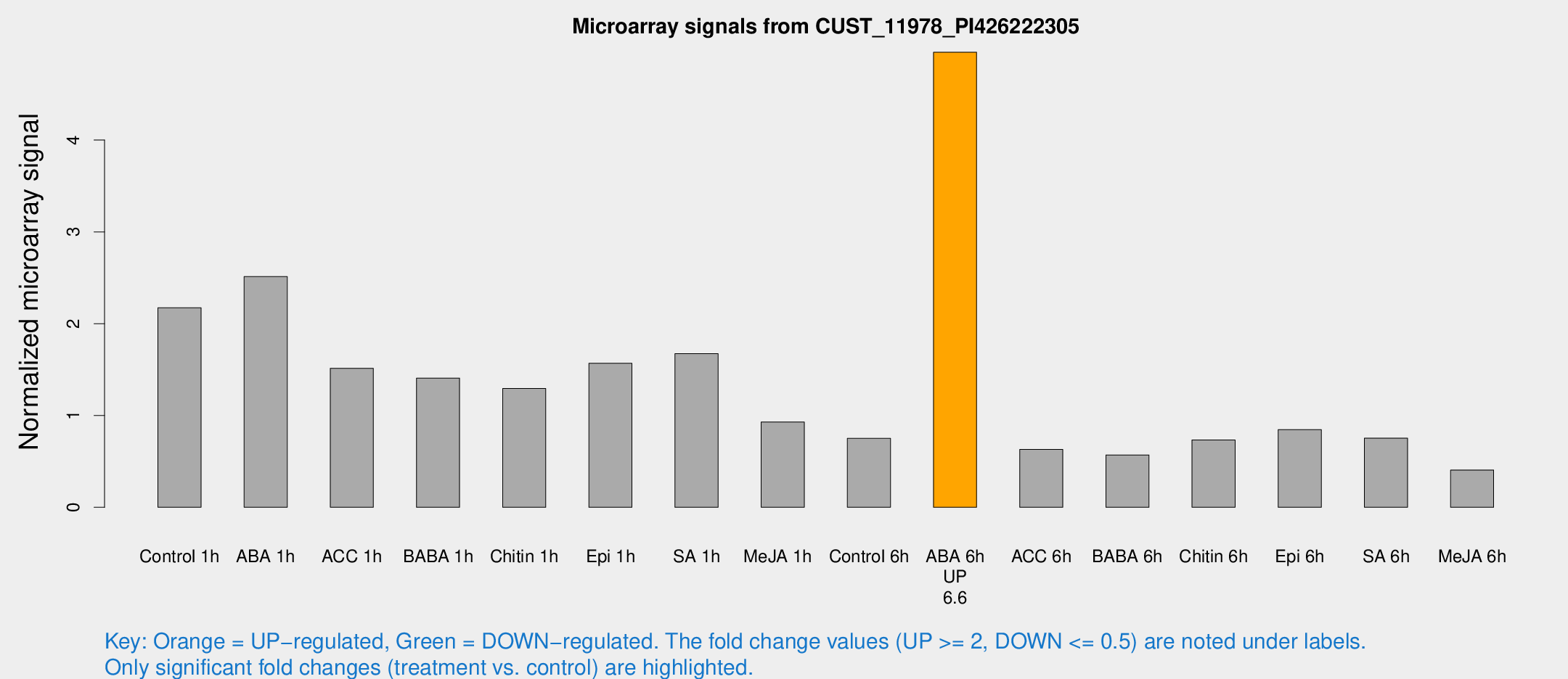
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 981.754 | 90.1013 | 2.17355 | 0.33971 |
| ABA 1h | 1206.84 | 545.921 | 2.51313 | 1.21706 |
| ACC 1h | 768.877 | 233.333 | 1.51473 | 0.412222 |
| BABA 1h | 646.219 | 143.793 | 1.40745 | 0.223404 |
| Chitin 1h | 584.614 | 207.922 | 1.29368 | 0.373173 |
| Epi 1h | 665.777 | 209.652 | 1.5687 | 0.519484 |
| SA 1h | 803.4 | 152.811 | 1.67293 | 0.2773 |
| Me-JA 1h | 339.999 | 19.911 | 0.929221 | 0.0634122 |
| Control 6h | 350.426 | 63.8222 | 0.750825 | 0.0845829 |
| ABA 6h | 2490.93 | 535.36 | 4.95705 | 0.972363 |
| ACC 6h | 327.179 | 33.7445 | 0.629844 | 0.160166 |
| BABA 6h | 287.893 | 24.0512 | 0.570286 | 0.0668419 |
| Chitin 6h | 352.34 | 32.5923 | 0.732969 | 0.0931482 |
| Epi 6h | 436.844 | 66.9283 | 0.84633 | 0.0682168 |
| SA 6h | 350.416 | 71.0268 | 0.753122 | 0.0858345 |
| Me-JA 6h | 204.701 | 75.0951 | 0.405771 | 0.125433 |
Source Transcript PGSC0003DMT400005956 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G79040.1 | +1 | 2e-25 | 96 | 81/145 (56%) | photosystem II subunit R | chr1:29736085-29736781 FORWARD LENGTH=140 |
