Probe CUST_11749_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11749_PI426222305 | JHI_St_60k_v1 | DMT400046721 | CAAAAACCACTTGCATATAAGTGGTCTCGACGAAGGGTTCATGAGAATGATGTTCGAAAA |
All Microarray Probes Designed to Gene DMG400018140
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11749_PI426222305 | JHI_St_60k_v1 | DMT400046721 | CAAAAACCACTTGCATATAAGTGGTCTCGACGAAGGGTTCATGAGAATGATGTTCGAAAA |
Microarray Signals from CUST_11749_PI426222305
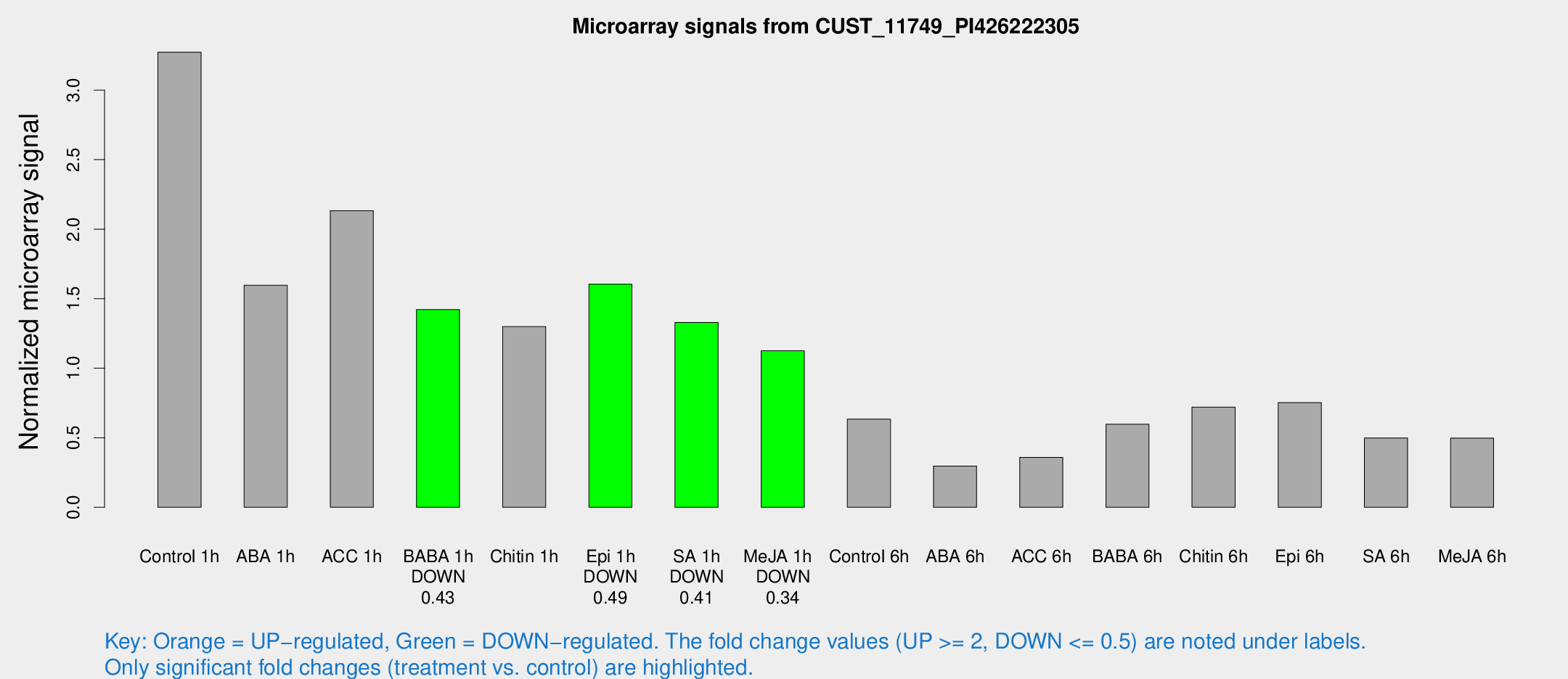
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 137.915 | 13.7329 | 3.2723 | 0.206052 |
| ABA 1h | 60.4261 | 9.12073 | 1.59624 | 0.363935 |
| ACC 1h | 93.5466 | 13.4257 | 2.13317 | 0.161843 |
| BABA 1h | 59.0488 | 9.93891 | 1.42129 | 0.125267 |
| Chitin 1h | 51.0488 | 11.4677 | 1.29927 | 0.302319 |
| Epi 1h | 58.3754 | 4.91478 | 1.60451 | 0.178831 |
| SA 1h | 57.7805 | 6.57556 | 1.32885 | 0.237368 |
| Me-JA 1h | 38.6103 | 4.14239 | 1.1264 | 0.120968 |
| Control 6h | 28.6824 | 8.18057 | 0.634465 | 0.130778 |
| ABA 6h | 14.9762 | 4.94375 | 0.297036 | 0.115109 |
| ACC 6h | 17.4893 | 4.41613 | 0.359253 | 0.0983179 |
| BABA 6h | 33.9118 | 11.7508 | 0.597187 | 0.319322 |
| Chitin 6h | 32.0777 | 4.27101 | 0.720572 | 0.0962345 |
| Epi 6h | 36.8126 | 7.49277 | 0.752824 | 0.142318 |
| SA 6h | 27.3813 | 14.8156 | 0.498831 | 0.445557 |
| Me-JA 6h | 22.0487 | 5.25246 | 0.497864 | 0.0965358 |
Source Transcript PGSC0003DMT400046721 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G48320.1 | +2 | 2e-168 | 491 | 252/468 (54%) | cytochrome P450, family 71, subfamily A, polypeptide 21 | chr3:17891241-17892804 FORWARD LENGTH=490 |
