Probe CUST_11052_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11052_PI426222305 | JHI_St_60k_v1 | DMT400078451 | CCGGAGGAAGGTGTCAAACAGATAATGCTACTAATAGATTTCTTTGTTCTGATACCAATG |
All Microarray Probes Designed to Gene DMG400030536
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11127_PI426222305 | JHI_St_60k_v1 | DMT400078452 | CCGCACACACTTGTGAGCATTTTTGAAAATTTTCTCTATGGATTATCCTGGCTTTAAATA |
| CUST_11052_PI426222305 | JHI_St_60k_v1 | DMT400078451 | CCGGAGGAAGGTGTCAAACAGATAATGCTACTAATAGATTTCTTTGTTCTGATACCAATG |
Microarray Signals from CUST_11052_PI426222305
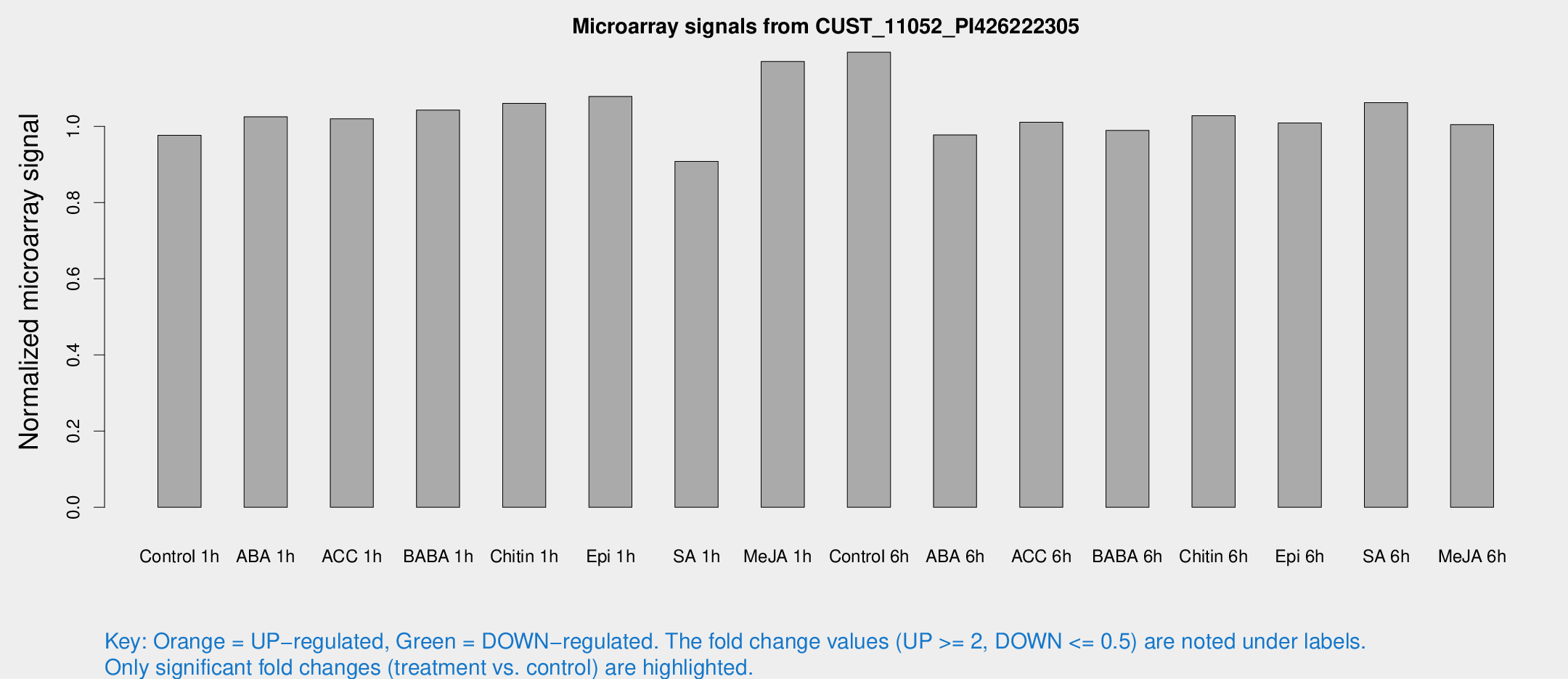
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.38985 | 3.50258 | 0.976524 | 0.538108 |
| ABA 1h | 5.92534 | 3.443 | 1.02506 | 0.594904 |
| ACC 1h | 6.84136 | 3.88986 | 1.01971 | 0.578258 |
| BABA 1h | 6.56471 | 3.80371 | 1.04293 | 0.604029 |
| Chitin 1h | 6.24165 | 3.62422 | 1.06041 | 0.614072 |
| Epi 1h | 6.10488 | 3.54309 | 1.07867 | 0.624918 |
| SA 1h | 6.09355 | 3.53409 | 0.908062 | 0.526229 |
| Me-JA 1h | 6.25439 | 3.63486 | 1.17052 | 0.678444 |
| Control 6h | 8.01909 | 3.76823 | 1.19464 | 0.597229 |
| ABA 6h | 6.79 | 3.93697 | 0.977356 | 0.56611 |
| ACC 6h | 7.75727 | 4.61331 | 1.01107 | 0.58641 |
| BABA 6h | 7.25376 | 4.21547 | 0.989277 | 0.572872 |
| Chitin 6h | 7.14498 | 4.13902 | 1.02823 | 0.595614 |
| Epi 6h | 7.49488 | 4.38009 | 1.00901 | 0.584271 |
| SA 6h | 6.86027 | 3.97534 | 1.06236 | 0.61554 |
| Me-JA 6h | 6.53734 | 3.79274 | 1.00462 | 0.582197 |
Source Transcript PGSC0003DMT400078451 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G25390.1 | +1 | 1e-159 | 481 | 294/640 (46%) | Protein kinase superfamily protein | chr1:8906640-8908800 REVERSE LENGTH=629 |
