Probe CUST_10838_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10838_PI426222305 | JHI_St_60k_v1 | DMT400032124 | TCGATTCCATGTGTCTTCTGAGGCAAGATGAGTCAATGAAGGCATTGTAAGCAGTAGCGA |
All Microarray Probes Designed to Gene DMG400012330
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10604_PI426222305 | JHI_St_60k_v1 | DMT400032123 | TCGATTCCATGTGTCTTCTGAGGCAAGATGAGTCAATGAAGGCATTGTAAGCAGTAGCGA |
| CUST_10838_PI426222305 | JHI_St_60k_v1 | DMT400032124 | TCGATTCCATGTGTCTTCTGAGGCAAGATGAGTCAATGAAGGCATTGTAAGCAGTAGCGA |
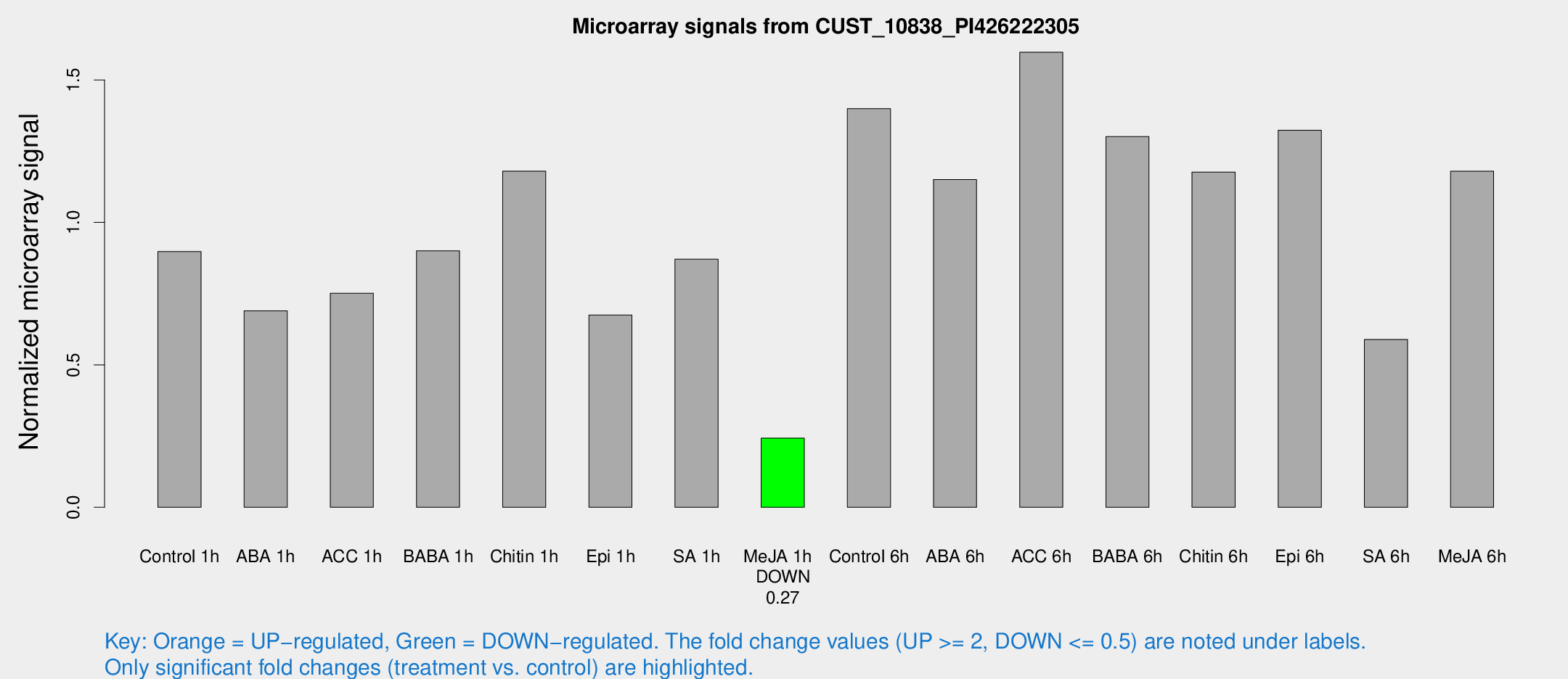
Microarray Signals from CUST_10838_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 24.3226 | 3.27483 | 0.897468 | 0.123739 |
| ABA 1h | 18.0459 | 5.34202 | 0.68964 | 0.225103 |
| ACC 1h | 22.2203 | 5.48306 | 0.751309 | 0.158189 |
| BABA 1h | 24.4948 | 5.29854 | 0.900203 | 0.316317 |
| Chitin 1h | 28.7731 | 3.41921 | 1.17979 | 0.151484 |
| Epi 1h | 16.2242 | 3.27055 | 0.674866 | 0.140904 |
| SA 1h | 24.4978 | 3.76826 | 0.870664 | 0.122659 |
| Me-JA 1h | 5.26155 | 3.02533 | 0.242975 | 0.137849 |
| Control 6h | 39.4146 | 8.38551 | 1.39901 | 0.190619 |
| ABA 6h | 33.7409 | 5.63193 | 1.15061 | 0.278928 |
| ACC 6h | 50.2236 | 7.48866 | 1.59719 | 0.149972 |
| BABA 6h | 41.9608 | 10.0278 | 1.30104 | 0.340768 |
| Chitin 6h | 36.6063 | 9.82435 | 1.17667 | 0.365042 |
| Epi 6h | 40.3361 | 4.23587 | 1.32369 | 0.2334 |
| SA 6h | 20.0868 | 8.95336 | 0.589131 | 0.292055 |
| Me-JA 6h | 31.4485 | 3.54542 | 1.17961 | 0.132937 |
Source Transcript PGSC0003DMT400032124 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G63850.1 | +3 | 0.0 | 658 | 359/527 (68%) | BTB/POZ domain-containing protein | chr1:23696962-23698708 FORWARD LENGTH=548 |
