Probe CUST_10784_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10784_PI426222305 | JHI_St_60k_v1 | DMT400031927 | TGGAAGGAAATCTAGCCCAAATTTTTTGGAGGGCACAATATTCTACTAGTGTTAATTGTT |
All Microarray Probes Designed to Gene DMG400012249
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10784_PI426222305 | JHI_St_60k_v1 | DMT400031927 | TGGAAGGAAATCTAGCCCAAATTTTTTGGAGGGCACAATATTCTACTAGTGTTAATTGTT |
| CUST_10617_PI426222305 | JHI_St_60k_v1 | DMT400031928 | GAAGCCAAGATCTCAATTTGGGATTTTCATCTGATTTCAAGACTATTACTGAGCTAATTC |
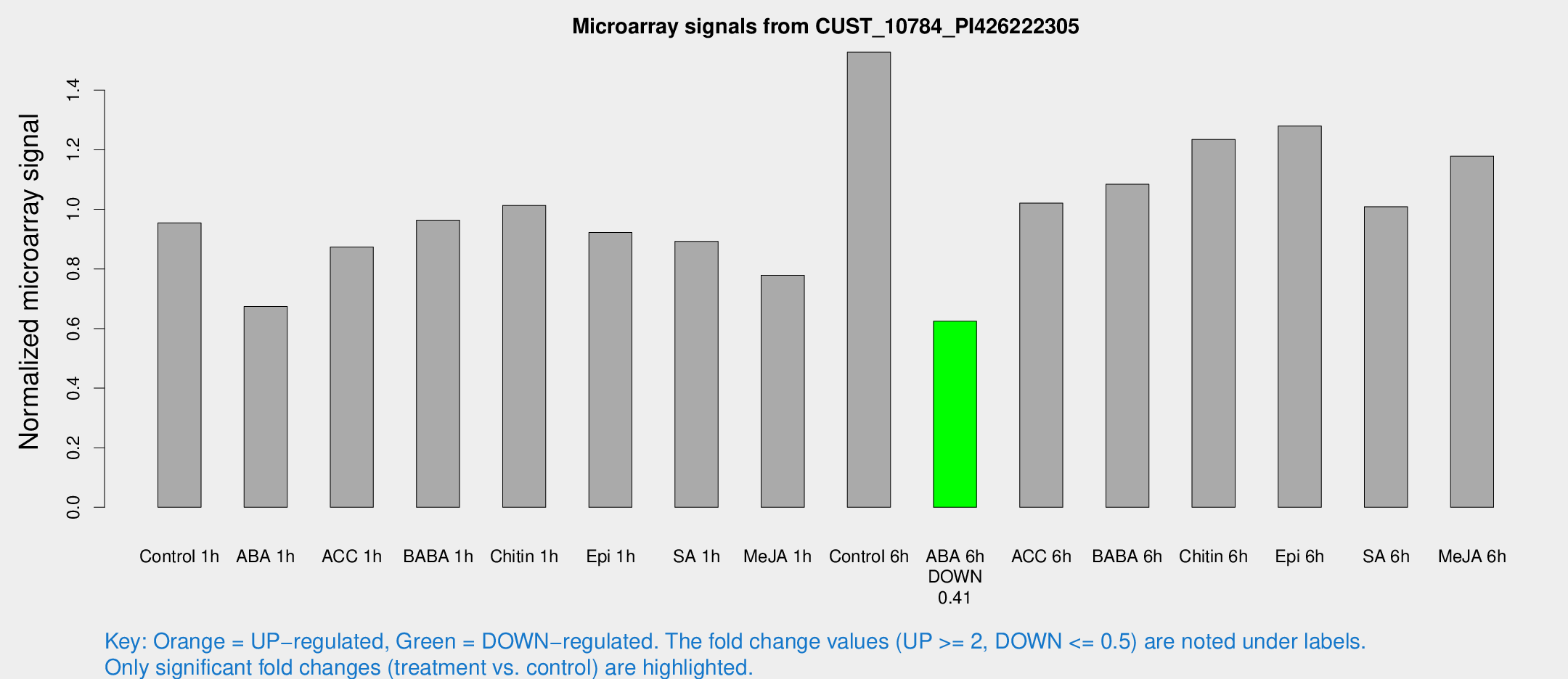
Microarray Signals from CUST_10784_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 84.209 | 25.2136 | 0.954616 | 0.230793 |
| ABA 1h | 48.7265 | 5.47099 | 0.673828 | 0.0646064 |
| ACC 1h | 77.5677 | 18.1215 | 0.873858 | 0.180037 |
| BABA 1h | 78.3507 | 15.8359 | 0.963753 | 0.118271 |
| Chitin 1h | 76.2704 | 13.5105 | 1.01332 | 0.14368 |
| Epi 1h | 64.4714 | 5.08271 | 0.922125 | 0.0731715 |
| SA 1h | 77.4796 | 15.8731 | 0.892662 | 0.169355 |
| Me-JA 1h | 51.8698 | 5.61608 | 0.778992 | 0.0721411 |
| Control 6h | 127.811 | 22.0162 | 1.52741 | 0.157297 |
| ABA 6h | 53.6698 | 5.01274 | 0.624432 | 0.0584468 |
| ACC 6h | 96.053 | 12.3678 | 1.0208 | 0.18176 |
| BABA 6h | 98.6825 | 8.63516 | 1.0844 | 0.0782889 |
| Chitin 6h | 106.552 | 7.72963 | 1.23466 | 0.0861374 |
| Epi 6h | 116.778 | 8.08371 | 1.27963 | 0.0883333 |
| SA 6h | 81.2931 | 7.8133 | 1.00888 | 0.0756879 |
| Me-JA 6h | 95.3772 | 8.20182 | 1.17888 | 0.0826367 |
Source Transcript PGSC0003DMT400031927 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G61850.4 | +2 | 6e-10 | 60 | 36/101 (36%) | Dof-type zinc finger DNA-binding family protein | chr3:22895495-22897150 FORWARD LENGTH=324 |
