Probe CUST_10538_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10538_PI426222305 | JHI_St_60k_v1 | DMT400031568 | CTGGTTAATGATTTTGTAGGTAAGGGCTATTTAAGGTTGTGTGGATCAAAGTCAATAGAA |
All Microarray Probes Designed to Gene DMG400012111
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10857_PI426222305 | JHI_St_60k_v1 | DMT400031569 | CTGGTTAATGATTTTGTAGGTAAGGGCTATTTAAGGTTGTGTGGATCAAAGTCAATAGAA |
| CUST_10538_PI426222305 | JHI_St_60k_v1 | DMT400031568 | CTGGTTAATGATTTTGTAGGTAAGGGCTATTTAAGGTTGTGTGGATCAAAGTCAATAGAA |
Microarray Signals from CUST_10538_PI426222305
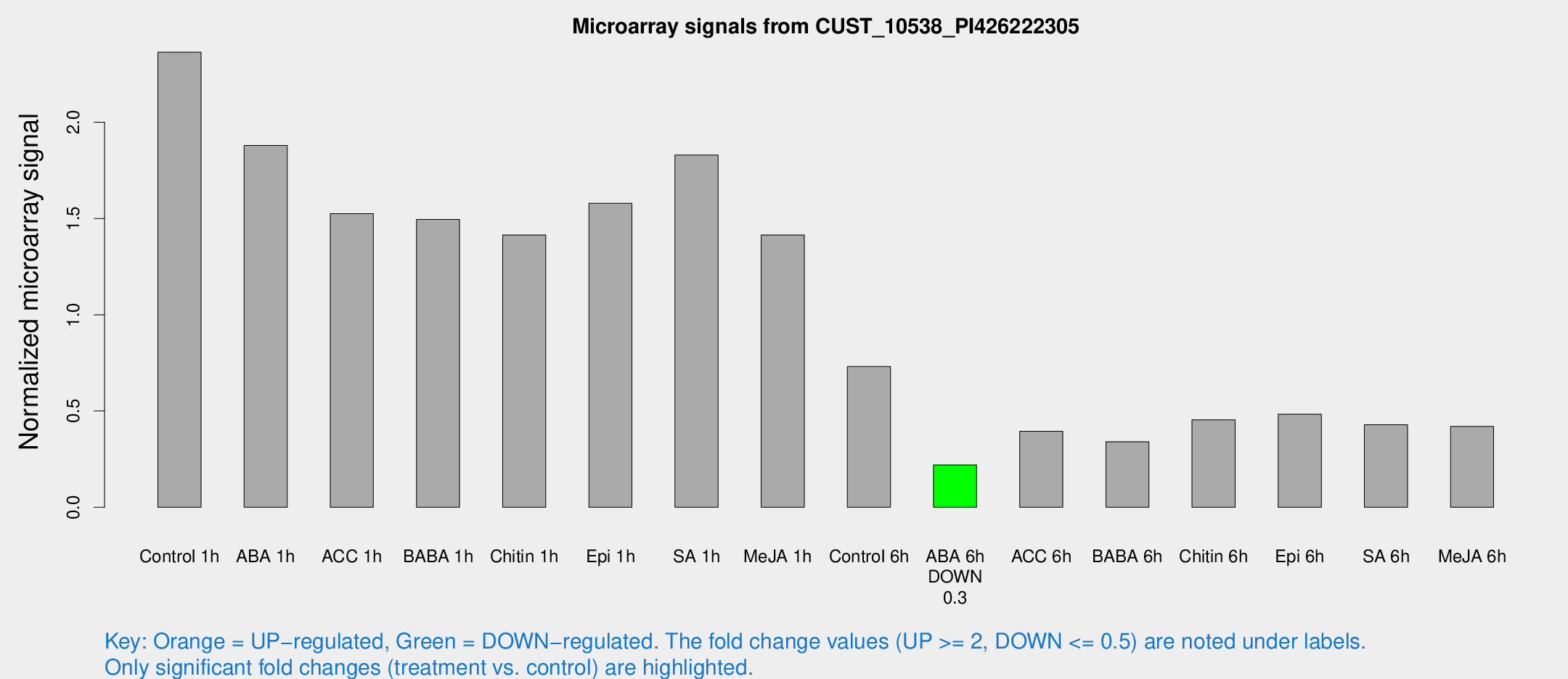
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 73173.2 | 4238.69 | 2.36332 | 0.193081 |
| ABA 1h | 51415.6 | 2976.48 | 1.87879 | 0.155121 |
| ACC 1h | 50445.8 | 9777.54 | 1.52525 | 0.219852 |
| BABA 1h | 46177.9 | 8519.83 | 1.49455 | 0.152815 |
| Chitin 1h | 39756.6 | 4563.4 | 1.41436 | 0.13441 |
| Epi 1h | 43328.6 | 6651.55 | 1.57912 | 0.248795 |
| SA 1h | 60022 | 11266.9 | 1.82956 | 0.313947 |
| Me-JA 1h | 35777.4 | 2328.63 | 1.41397 | 0.0816358 |
| Control 6h | 22937.3 | 2728.72 | 0.731179 | 0.0543449 |
| ABA 6h | 7361.05 | 971.331 | 0.220183 | 0.0198825 |
| ACC 6h | 14086 | 1294.48 | 0.394753 | 0.0735316 |
| BABA 6h | 12970.8 | 3676.33 | 0.340761 | 0.130743 |
| Chitin 6h | 14982.3 | 911.146 | 0.454504 | 0.0262412 |
| Epi 6h | 16851.5 | 974.73 | 0.48345 | 0.0407029 |
| SA 6h | 13730.3 | 2725.09 | 0.42894 | 0.0470771 |
| Me-JA 6h | 13014.7 | 1171.75 | 0.420278 | 0.0302253 |
Source Transcript PGSC0003DMT400031568 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G32900.1 | +2 | 0.0 | 876 | 444/611 (73%) | UDP-Glycosyltransferase superfamily protein | chr1:11920582-11923506 REVERSE LENGTH=610 |
