Probe CUST_10473_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10473_PI426222305 | JHI_St_60k_v1 | DMT400031995 | ACGTCCTACCTTTAAATTCTGAATGCGCCTCTAAACACTCCGGTTTATGTTGCTTGCAGG |
All Microarray Probes Designed to Gene DMG400012274
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10473_PI426222305 | JHI_St_60k_v1 | DMT400031995 | ACGTCCTACCTTTAAATTCTGAATGCGCCTCTAAACACTCCGGTTTATGTTGCTTGCAGG |
| CUST_10595_PI426222305 | JHI_St_60k_v1 | DMT400031994 | GATGCTGAAATTGTATATGCTGCAACAGAGAAAGCAGAACGCGAATTGAAGTGGAAGGGG |
Microarray Signals from CUST_10473_PI426222305
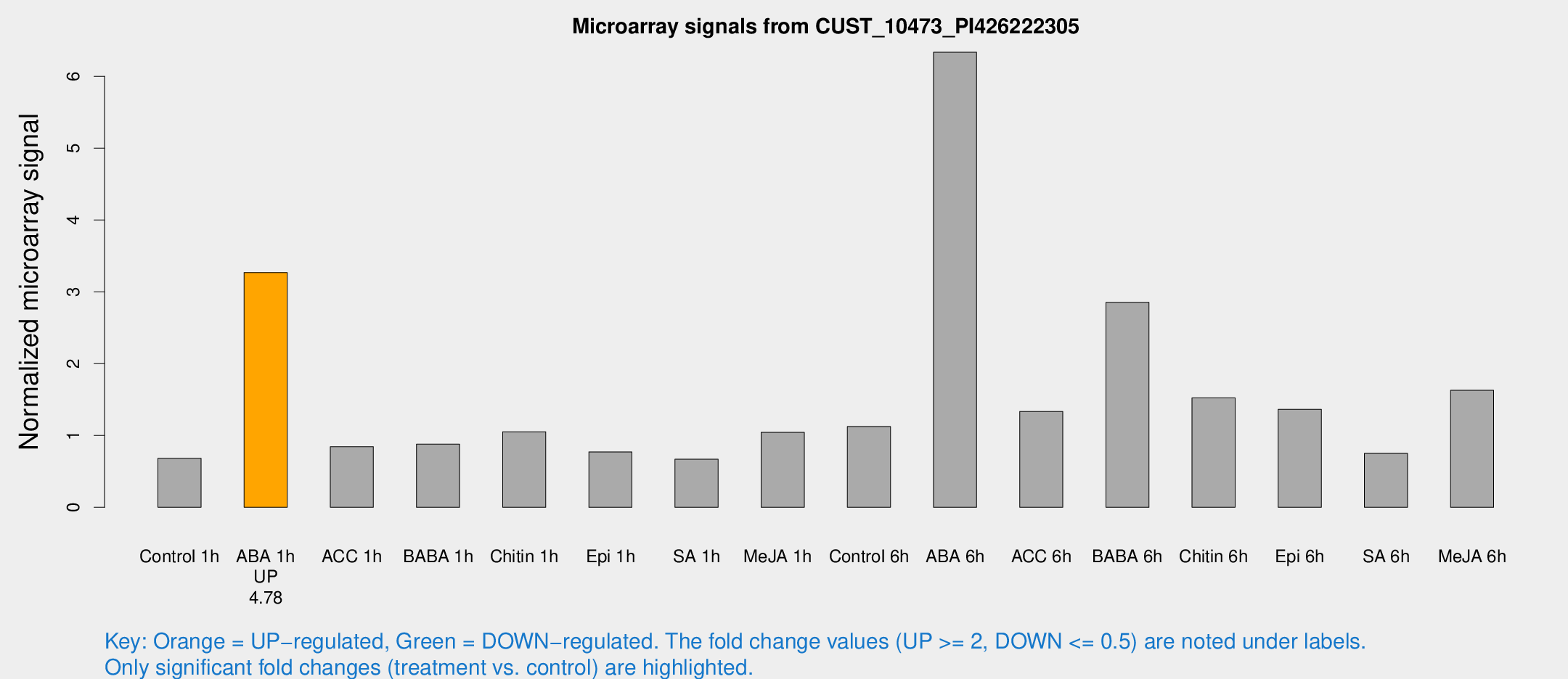
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.67948 | 3.2918 | 0.683406 | 0.395826 |
| ABA 1h | 24.9873 | 4.7538 | 3.26896 | 0.497572 |
| ACC 1h | 7.29478 | 3.88004 | 0.843365 | 0.447511 |
| BABA 1h | 7.10394 | 3.44942 | 0.878206 | 0.437128 |
| Chitin 1h | 8.06817 | 3.28162 | 1.05236 | 0.44913 |
| Epi 1h | 5.54808 | 3.22662 | 0.770653 | 0.447518 |
| SA 1h | 5.71347 | 3.24901 | 0.669265 | 0.380672 |
| Me-JA 1h | 7.62203 | 3.36382 | 1.04442 | 0.516404 |
| Control 6h | 11.7767 | 5.86589 | 1.12501 | 0.570759 |
| ABA 6h | 60.7782 | 15.249 | 6.33543 | 1.74742 |
| ACC 6h | 14.6377 | 4.58911 | 1.33489 | 0.598583 |
| BABA 6h | 27.7158 | 5.32003 | 2.85269 | 0.525624 |
| Chitin 6h | 13.6265 | 3.90286 | 1.52429 | 0.449342 |
| Epi 6h | 14.5896 | 4.50081 | 1.36523 | 0.498866 |
| SA 6h | 6.17289 | 3.57238 | 0.751116 | 0.4345 |
| Me-JA 6h | 13.8039 | 3.47414 | 1.62813 | 0.428523 |
Source Transcript PGSC0003DMT400031995 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G23920.1 | +1 | 7e-178 | 510 | 261/318 (82%) | UDP-D-glucose/UDP-D-galactose 4-epimerase 2 | chr4:12431416-12433666 FORWARD LENGTH=350 |
