Probe CUST_10186_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10186_PI426222305 | JHI_St_60k_v1 | DMT400070675 | GTTGTCGTGTTCAAAACATTATTGTTCCTCGTCCTTCACCTCCAAAGCTTTTATCTTAAA |
All Microarray Probes Designed to Gene DMG400027473
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10198_PI426222305 | JHI_St_60k_v1 | DMT400070674 | TCCAATGGAAATATTCAACAAGGTTGTAGAGATCCAGAAACACGTGATTGGAACAAAGTG |
| CUST_10186_PI426222305 | JHI_St_60k_v1 | DMT400070675 | GTTGTCGTGTTCAAAACATTATTGTTCCTCGTCCTTCACCTCCAAAGCTTTTATCTTAAA |
Microarray Signals from CUST_10186_PI426222305
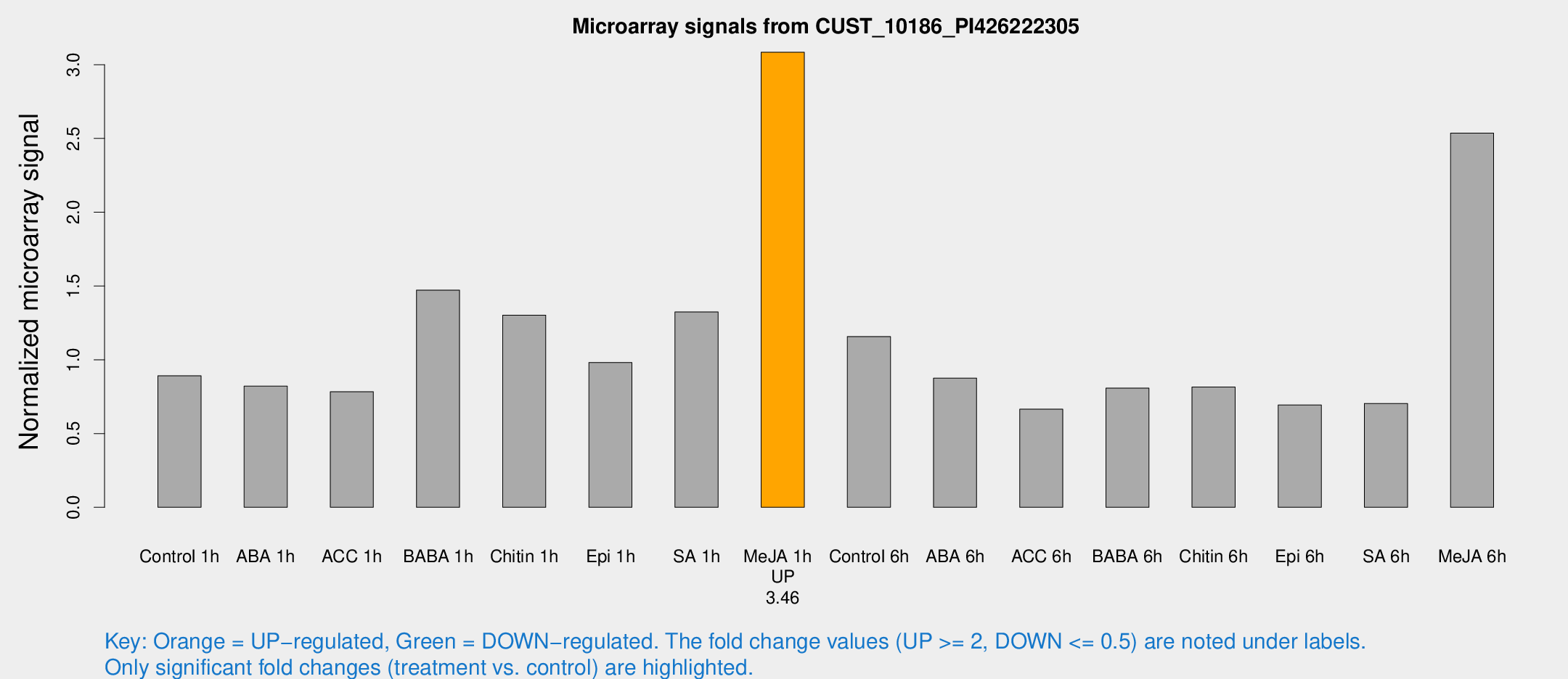
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1810.5 | 439.016 | 0.891439 | 0.273156 |
| ABA 1h | 1476.82 | 388.165 | 0.82132 | 0.155556 |
| ACC 1h | 1679.94 | 491.925 | 0.782919 | 0.199835 |
| BABA 1h | 2736.3 | 261.183 | 1.47229 | 0.0850201 |
| Chitin 1h | 2251.67 | 183.102 | 1.30238 | 0.172637 |
| Epi 1h | 1744.26 | 447.745 | 0.981437 | 0.280228 |
| SA 1h | 2737.33 | 634.925 | 1.32412 | 0.357132 |
| Me-JA 1h | 4857.39 | 503.518 | 3.08437 | 0.178087 |
| Control 6h | 2304.66 | 453.764 | 1.15754 | 0.134766 |
| ABA 6h | 1785.61 | 115.856 | 0.875736 | 0.0831505 |
| ACC 6h | 1470.91 | 139.633 | 0.665011 | 0.0859178 |
| BABA 6h | 1750.55 | 200.409 | 0.807351 | 0.129463 |
| Chitin 6h | 1707.96 | 292.945 | 0.815276 | 0.163236 |
| Epi 6h | 1576.47 | 330.044 | 0.693684 | 0.229861 |
| SA 6h | 1390.19 | 269.13 | 0.703969 | 0.0713104 |
| Me-JA 6h | 4868.72 | 464.213 | 2.53601 | 0.310912 |
Source Transcript PGSC0003DMT400070675 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G14220.1 | +2 | 5e-12 | 64 | 55/196 (28%) | Ribonuclease T2 family protein | chr1:4858642-4859601 REVERSE LENGTH=228 |
