Probe CUST_10050_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10050_PI426222305 | JHI_St_60k_v1 | DMT400028158 | GGCAAAGGAATACCTAATAGTGTGTCAATATAGTACTTTCATTTAAATCTCCACATCGTC |
All Microarray Probes Designed to Gene DMG400010859
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10030_PI426222305 | JHI_St_60k_v1 | DMT400028157 | GGCAAAGGAATACCTAATAGTGTGTCAATATAGTACTTTCATTTAAATCTCCACATCGTC |
| CUST_10050_PI426222305 | JHI_St_60k_v1 | DMT400028158 | GGCAAAGGAATACCTAATAGTGTGTCAATATAGTACTTTCATTTAAATCTCCACATCGTC |
Microarray Signals from CUST_10050_PI426222305
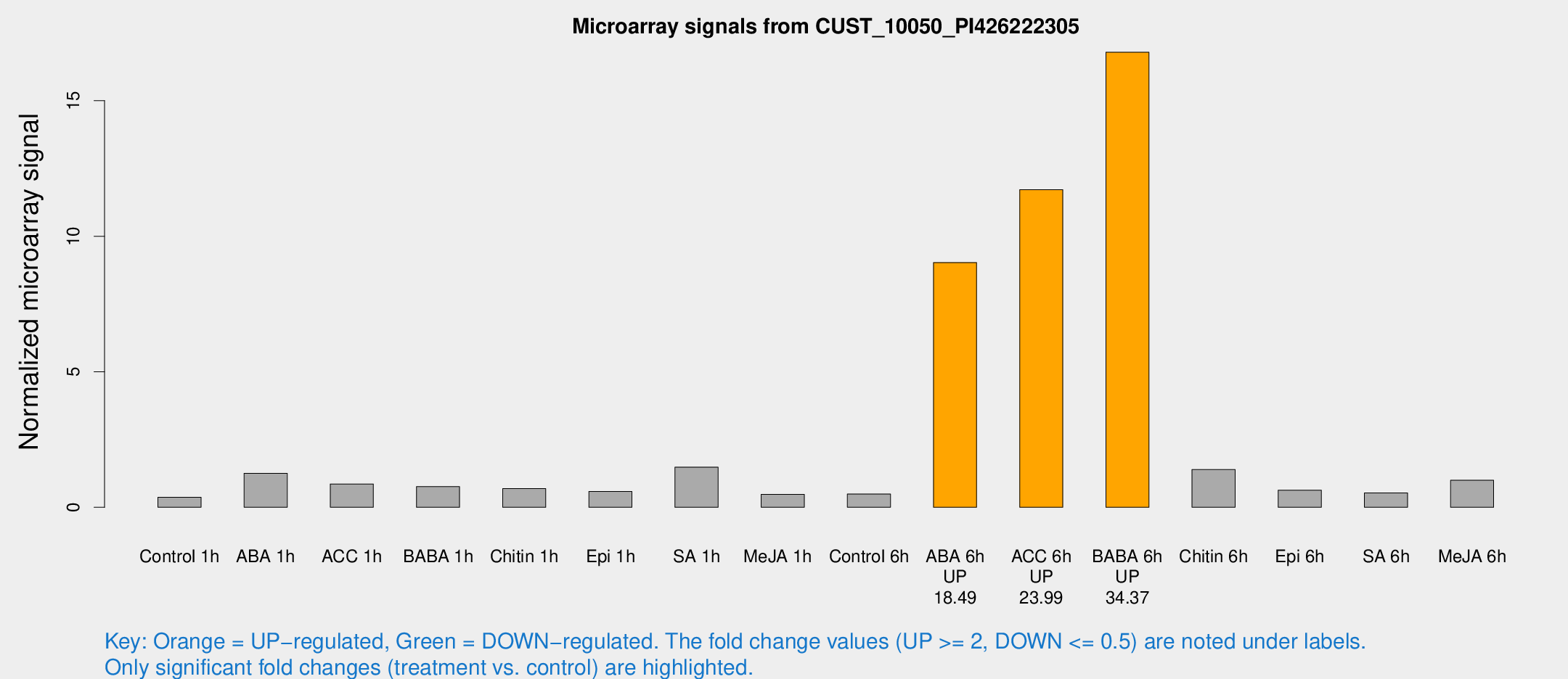
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 16.2696 | 9.67262 | 0.372301 | 0.241445 |
| ABA 1h | 37.9021 | 12.8226 | 1.25602 | 0.491851 |
| ACC 1h | 29.6737 | 7.75884 | 0.861657 | 0.220803 |
| BABA 1h | 32.3595 | 13.9347 | 0.76146 | 0.614581 |
| Chitin 1h | 20.1802 | 4.95202 | 0.688916 | 0.161527 |
| Epi 1h | 15.8659 | 3.20451 | 0.584551 | 0.127794 |
| SA 1h | 57.8695 | 28.0174 | 1.48217 | 0.658666 |
| Me-JA 1h | 12.6283 | 3.31961 | 0.479395 | 0.144905 |
| Control 6h | 15.6183 | 3.31483 | 0.488377 | 0.139861 |
| ABA 6h | 331.899 | 107.52 | 9.02937 | 3.00171 |
| ACC 6h | 411.744 | 24.1252 | 11.7182 | 1.98097 |
| BABA 6h | 689.865 | 279.021 | 16.7872 | 8.32012 |
| Chitin 6h | 56.5829 | 26.2189 | 1.39039 | 0.717628 |
| Epi 6h | 30.0939 | 12.2158 | 0.628037 | 0.505234 |
| SA 6h | 27.7184 | 19.6361 | 0.530992 | 0.491747 |
| Me-JA 6h | 34.8298 | 12.871 | 0.996625 | 0.304671 |
Source Transcript PGSC0003DMT400028158 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G55020.1 | +3 | 0.0 | 1242 | 591/852 (69%) | lipoxygenase 1 | chr1:20525798-20530143 FORWARD LENGTH=859 |
