Probe CUST_9802_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9802_PI426222305 | JHI_St_60k_v1 | DMT400016584 | CGCAAAATTGGTCTCAAAATAATGGCTTCCATCATAATTTTATGAGACCTTCTTGACCAT |
All Microarray Probes Designed to Gene DMG400006482
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9770_PI426222305 | JHI_St_60k_v1 | DMT400016585 | CGCAAAATTGGTCTCAAAATAATGGCTTCCATCATAATTTTATGAGACCTTCTTGACCAT |
| CUST_9802_PI426222305 | JHI_St_60k_v1 | DMT400016584 | CGCAAAATTGGTCTCAAAATAATGGCTTCCATCATAATTTTATGAGACCTTCTTGACCAT |
Microarray Signals from CUST_9802_PI426222305
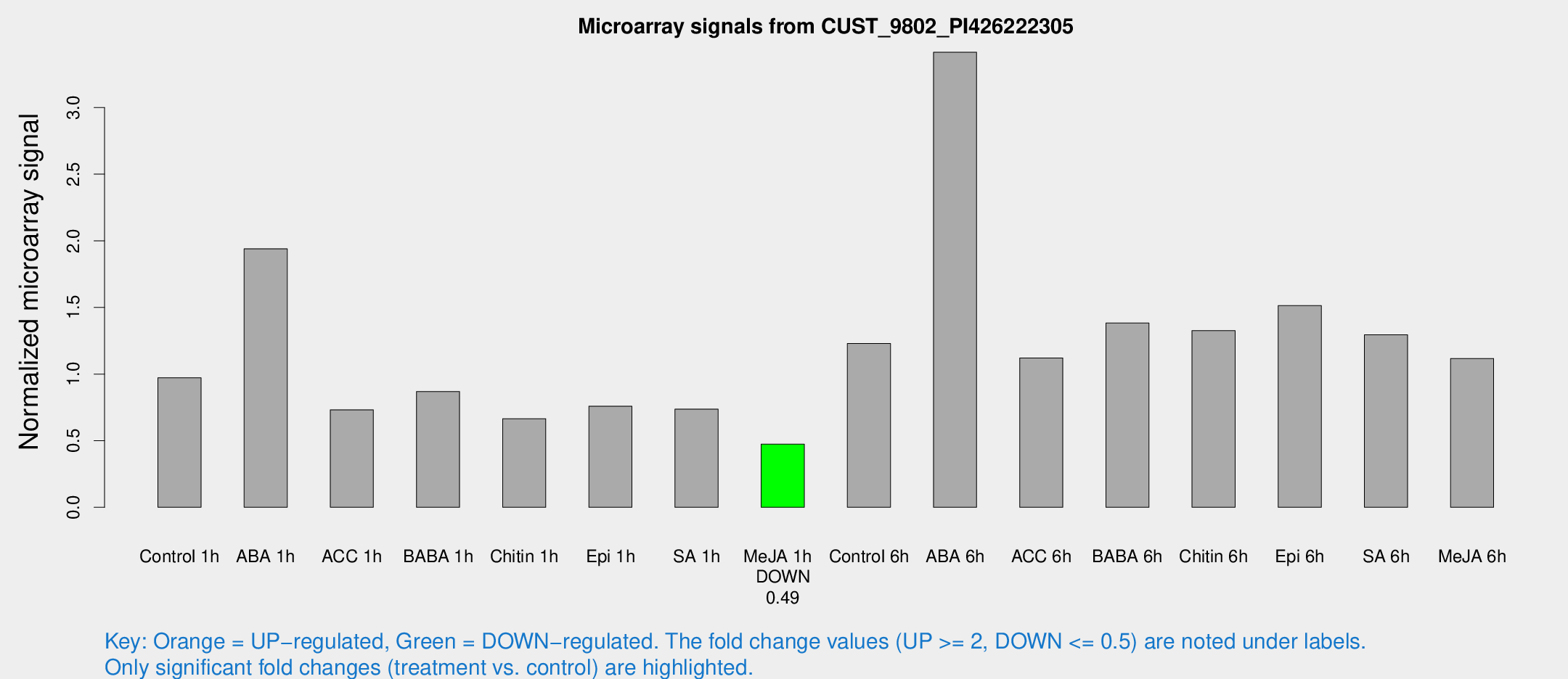
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 120.648 | 17.006 | 0.972355 | 0.0895913 |
| ABA 1h | 214.17 | 31.8029 | 1.93998 | 0.238702 |
| ACC 1h | 102.7 | 29.369 | 0.73188 | 0.223624 |
| BABA 1h | 105.022 | 17.2361 | 0.868713 | 0.0690973 |
| Chitin 1h | 73.769 | 8.54544 | 0.665175 | 0.0499966 |
| Epi 1h | 83.2447 | 15.313 | 0.758628 | 0.149529 |
| SA 1h | 95.5675 | 16.6257 | 0.73718 | 0.156625 |
| Me-JA 1h | 47.5063 | 4.44202 | 0.473763 | 0.0445637 |
| Control 6h | 170.921 | 51.9967 | 1.2293 | 0.385535 |
| ABA 6h | 452.116 | 63.869 | 3.41504 | 0.359936 |
| ACC 6h | 157.361 | 11.0271 | 1.12103 | 0.0798387 |
| BABA 6h | 189.184 | 11.7122 | 1.38299 | 0.0854304 |
| Chitin 6h | 172.033 | 10.7552 | 1.3259 | 0.0829037 |
| Epi 6h | 212.602 | 30.0537 | 1.51367 | 0.344766 |
| SA 6h | 157.371 | 14.8634 | 1.29478 | 0.200776 |
| Me-JA 6h | 149.189 | 46.5502 | 1.11681 | 0.263693 |
Source Transcript PGSC0003DMT400016584 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G36920.1 | +2 | 2e-94 | 254 | 171/323 (53%) | Integrase-type DNA-binding superfamily protein | chr4:17400998-17403140 FORWARD LENGTH=432 |
