Probe CUST_9752_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9752_PI426222305 | JHI_St_60k_v1 | DMT400038534 | TTTGTCAGTGTCACTGATAGCGAGGCTTTTTCAGAATTGAGGATCGATGAAACCAGATGA |
All Microarray Probes Designed to Gene DMG400014875
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9783_PI426222305 | JHI_St_60k_v1 | DMT400038535 | CTTTAATTTCTGCATTGGATAATAGGCAAATGCCTCTTGACAATGCATCTGTATTGATAG |
| CUST_9752_PI426222305 | JHI_St_60k_v1 | DMT400038534 | TTTGTCAGTGTCACTGATAGCGAGGCTTTTTCAGAATTGAGGATCGATGAAACCAGATGA |
Microarray Signals from CUST_9752_PI426222305
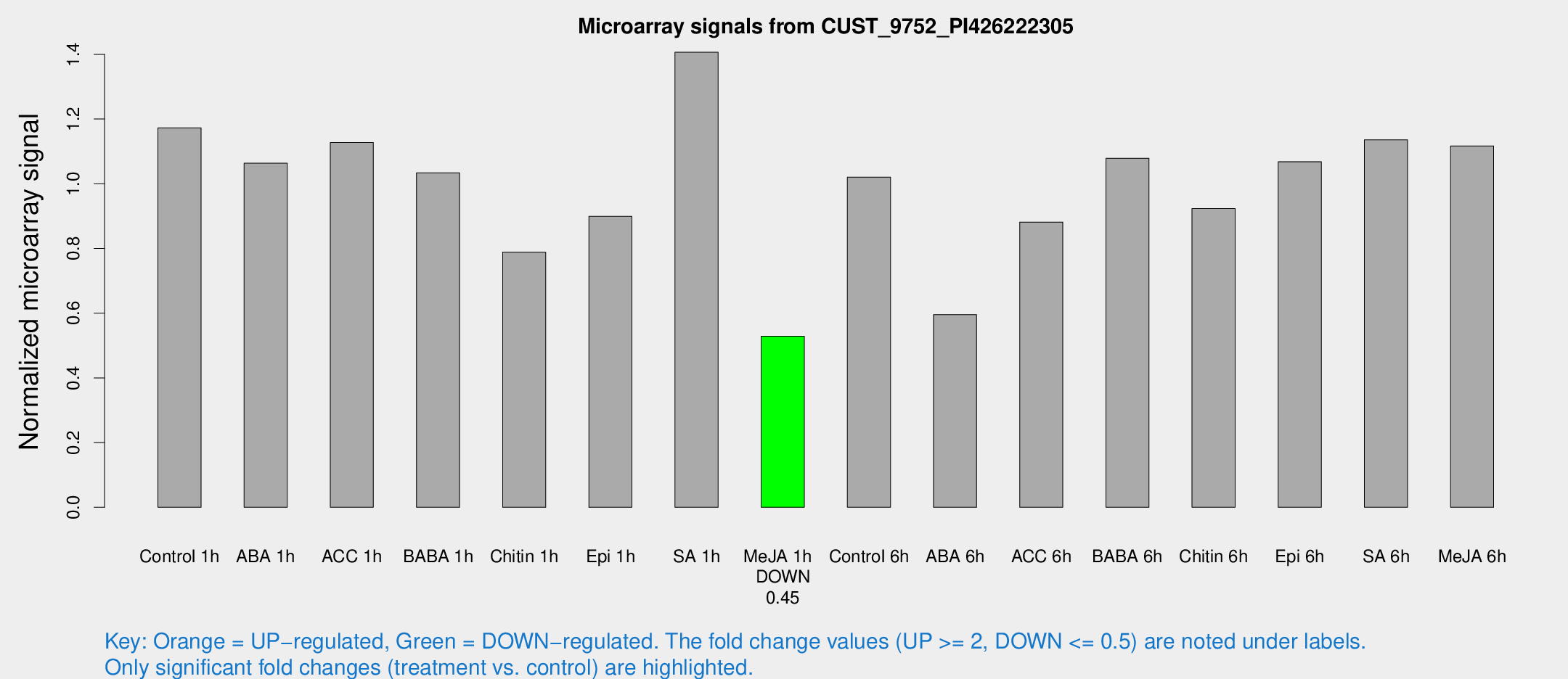
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3341.8 | 547.148 | 1.17266 | 0.110303 |
| ABA 1h | 2621.98 | 151.895 | 1.0637 | 0.0614262 |
| ACC 1h | 3347.96 | 626.82 | 1.12723 | 0.146134 |
| BABA 1h | 2822.31 | 388.456 | 1.03396 | 0.0691323 |
| Chitin 1h | 2001.32 | 243.98 | 0.788544 | 0.0730255 |
| Epi 1h | 2166.9 | 125.433 | 0.899186 | 0.051932 |
| SA 1h | 4045.88 | 364.057 | 1.4065 | 0.0812127 |
| Me-JA 1h | 1200.58 | 69.4946 | 0.528656 | 0.0305576 |
| Control 6h | 2954.8 | 538.266 | 1.02025 | 0.115223 |
| ABA 6h | 1770.48 | 144.047 | 0.595605 | 0.0344089 |
| ACC 6h | 2831.12 | 255.896 | 0.881555 | 0.050912 |
| BABA 6h | 3366.47 | 195.048 | 1.07887 | 0.0623009 |
| Chitin 6h | 2735.25 | 158.252 | 0.923375 | 0.0533268 |
| Epi 6h | 3354.15 | 194.208 | 1.06813 | 0.0914556 |
| SA 6h | 3274.64 | 679.716 | 1.13555 | 0.132991 |
| Me-JA 6h | 3243.63 | 734.16 | 1.11694 | 0.157602 |
Source Transcript PGSC0003DMT400038534 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G24010.1 | +1 | 0.0 | 548 | 347/740 (47%) | Protein kinase superfamily protein | chr5:8113910-8116384 FORWARD LENGTH=824 |
