Probe CUST_9722_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9722_PI426222305 | JHI_St_60k_v1 | DMT400008750 | ATCATTGTCTTAGTTACCGGTACATACACATCCATGGAGGAAATAATACACCTCATGTAA |
All Microarray Probes Designed to Gene DMG400003397
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9693_PI426222305 | JHI_St_60k_v1 | DMT400008751 | CCGGTACATACACATCCATGGAGGAAATAATACACCTCATGTAATTATGTATTTTACTGA |
| CUST_9722_PI426222305 | JHI_St_60k_v1 | DMT400008750 | ATCATTGTCTTAGTTACCGGTACATACACATCCATGGAGGAAATAATACACCTCATGTAA |
Microarray Signals from CUST_9722_PI426222305
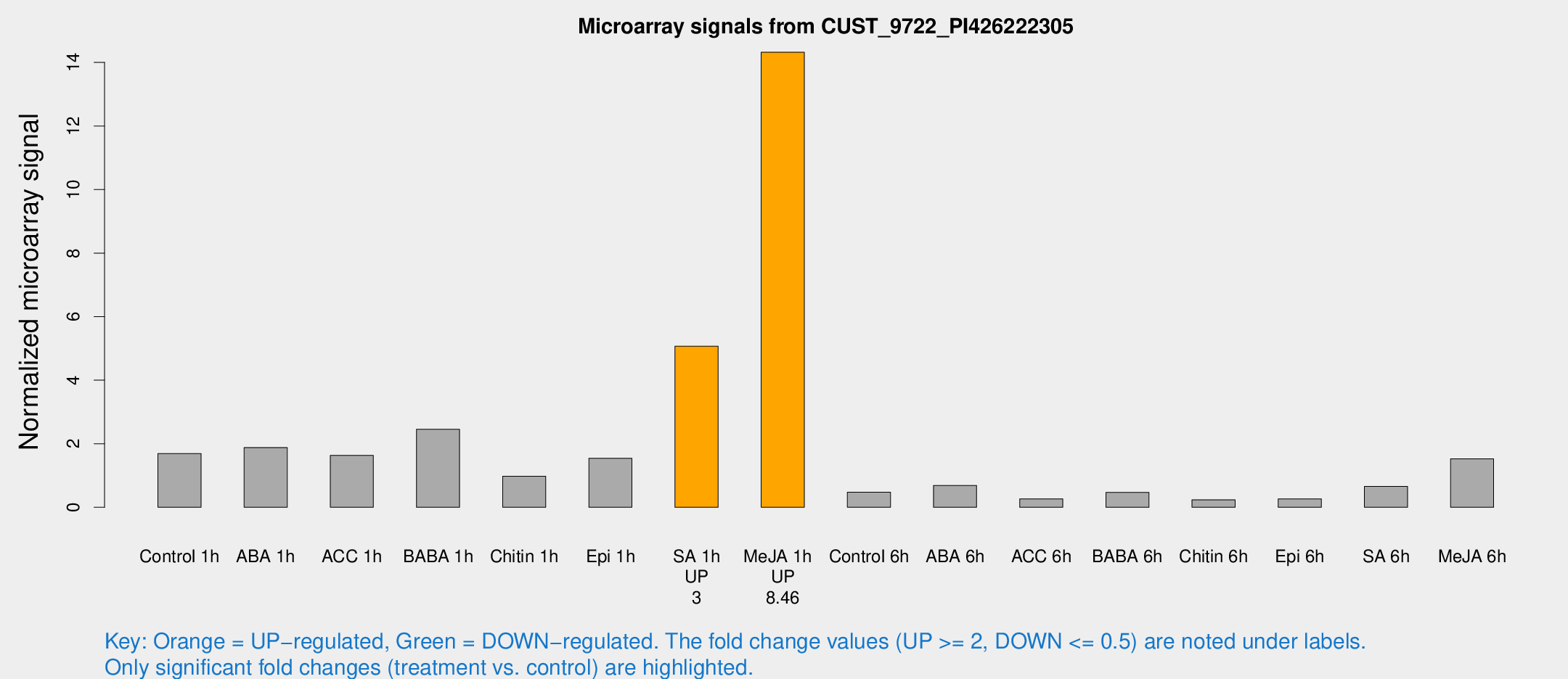
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 4168.97 | 750.276 | 1.6919 | 0.235832 |
| ABA 1h | 4965.55 | 2318.39 | 1.88081 | 0.856998 |
| ACC 1h | 5126.17 | 2631.57 | 1.63593 | 0.864235 |
| BABA 1h | 5772.68 | 788.511 | 2.45482 | 0.141737 |
| Chitin 1h | 2127.67 | 211.342 | 0.979997 | 0.0636556 |
| Epi 1h | 3248.65 | 414.785 | 1.54299 | 0.195322 |
| SA 1h | 13371.2 | 3786.09 | 5.06815 | 1.42631 |
| Me-JA 1h | 28427.4 | 3828.13 | 14.321 | 1.47156 |
| Control 6h | 1336.51 | 549.246 | 0.473591 | 0.165467 |
| ABA 6h | 1880.66 | 498.186 | 0.688235 | 0.155312 |
| ACC 6h | 1084.6 | 580.672 | 0.265363 | 0.286664 |
| BABA 6h | 1338.49 | 324.657 | 0.46752 | 0.133275 |
| Chitin 6h | 602.786 | 62.6254 | 0.234328 | 0.0318687 |
| Epi 6h | 739.362 | 158.898 | 0.263057 | 0.0593797 |
| SA 6h | 1660.9 | 381.11 | 0.658937 | 0.0992568 |
| Me-JA 6h | 3640.79 | 233.969 | 1.52378 | 0.174858 |
Source Transcript PGSC0003DMT400008750 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G28960.1 | +1 | 4e-139 | 405 | 201/352 (57%) | Transmembrane amino acid transporter family protein | chr3:10984245-10985767 REVERSE LENGTH=405 |
