Probe CUST_9550_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9550_PI426222305 | JHI_St_60k_v1 | DMT400089692 | TGTGTGGATCTATGGTGGAAAGGGCAGCCAATTTCATTCTGCTCATTATGGAAGATTTAG |
All Microarray Probes Designed to Gene DMG400039263
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9550_PI426222305 | JHI_St_60k_v1 | DMT400089692 | TGTGTGGATCTATGGTGGAAAGGGCAGCCAATTTCATTCTGCTCATTATGGAAGATTTAG |
Microarray Signals from CUST_9550_PI426222305
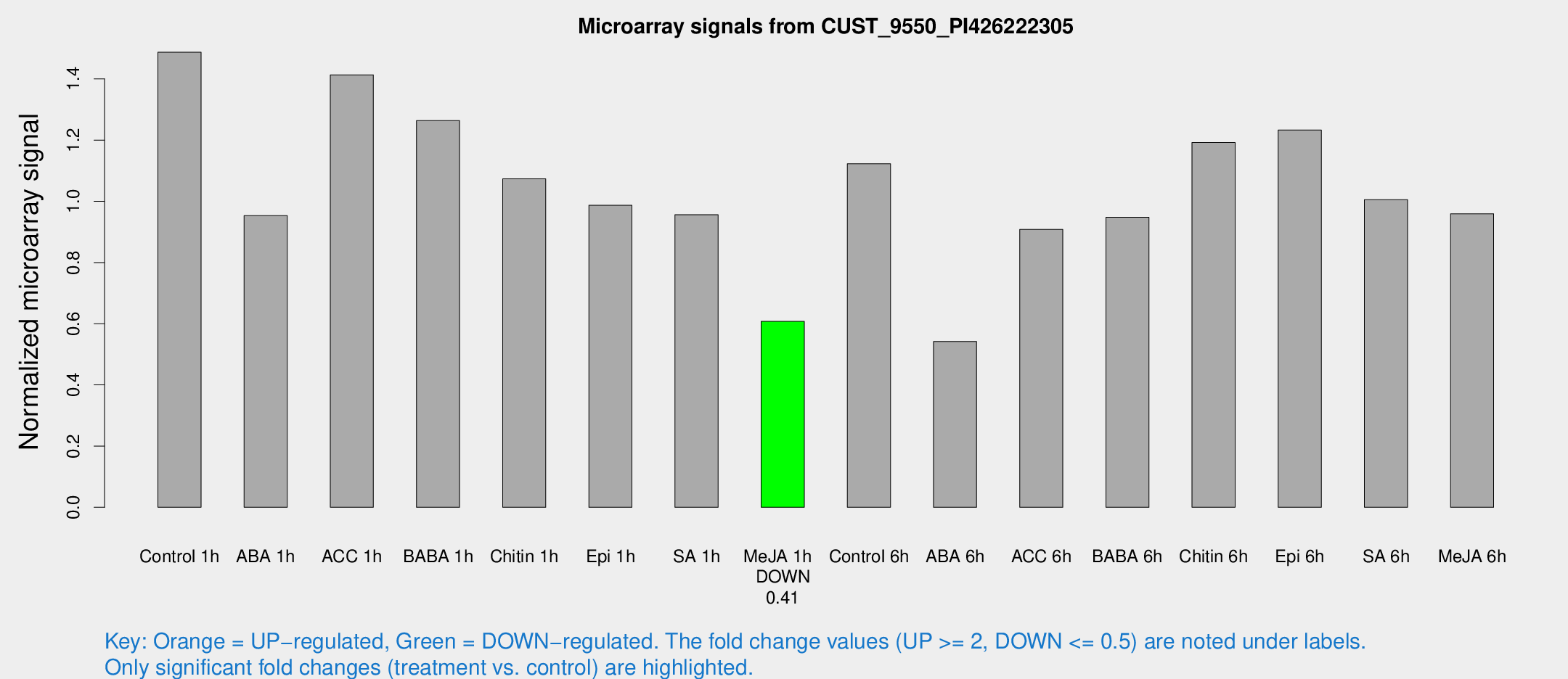
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 363.766 | 58.2111 | 1.48721 | 0.135678 |
| ABA 1h | 201.586 | 11.9933 | 0.953115 | 0.0664141 |
| ACC 1h | 360.884 | 67.0269 | 1.41292 | 0.185521 |
| BABA 1h | 299.858 | 48.835 | 1.2639 | 0.0951759 |
| Chitin 1h | 238.713 | 43.7497 | 1.07335 | 0.174358 |
| Epi 1h | 205.94 | 20.3534 | 0.98707 | 0.113767 |
| SA 1h | 274.358 | 88.3437 | 0.955905 | 0.406046 |
| Me-JA 1h | 119.934 | 13.0026 | 0.607405 | 0.0384281 |
| Control 6h | 281.857 | 61.6875 | 1.12251 | 0.156046 |
| ABA 6h | 138.062 | 8.5953 | 0.541808 | 0.0497913 |
| ACC 6h | 250.837 | 23.5839 | 0.908032 | 0.0539876 |
| BABA 6h | 257.528 | 32.2245 | 0.947721 | 0.147458 |
| Chitin 6h | 311.926 | 49.6223 | 1.19198 | 0.253506 |
| Epi 6h | 339.373 | 52.0901 | 1.23277 | 0.218184 |
| SA 6h | 254.984 | 65.1046 | 1.00519 | 0.1775 |
| Me-JA 6h | 235.408 | 42.5621 | 0.959136 | 0.0932283 |
Source Transcript PGSC0003DMT400089692 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G13840.1 | +1 | 2e-150 | 445 | 261/558 (47%) | GRAS family transcription factor | chr3:4555305-4556837 REVERSE LENGTH=510 |
