Probe CUST_9528_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9528_PI426222305 | JHI_St_60k_v1 | DMT400023617 | GCTATCCAAAATGCCTTATTCTAAGAATGAGAACAGAACTTCTATATGGTTGGAAGAGAT |
All Microarray Probes Designed to Gene DMG400009144
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9165_PI426222305 | JHI_St_60k_v1 | DMT400023618 | GGACTGTCGCTTTAGTATATGGATTCTTTAAACAGAACTTCTATATGGTTGGAAGAGATA |
| CUST_9528_PI426222305 | JHI_St_60k_v1 | DMT400023617 | GCTATCCAAAATGCCTTATTCTAAGAATGAGAACAGAACTTCTATATGGTTGGAAGAGAT |
Microarray Signals from CUST_9528_PI426222305
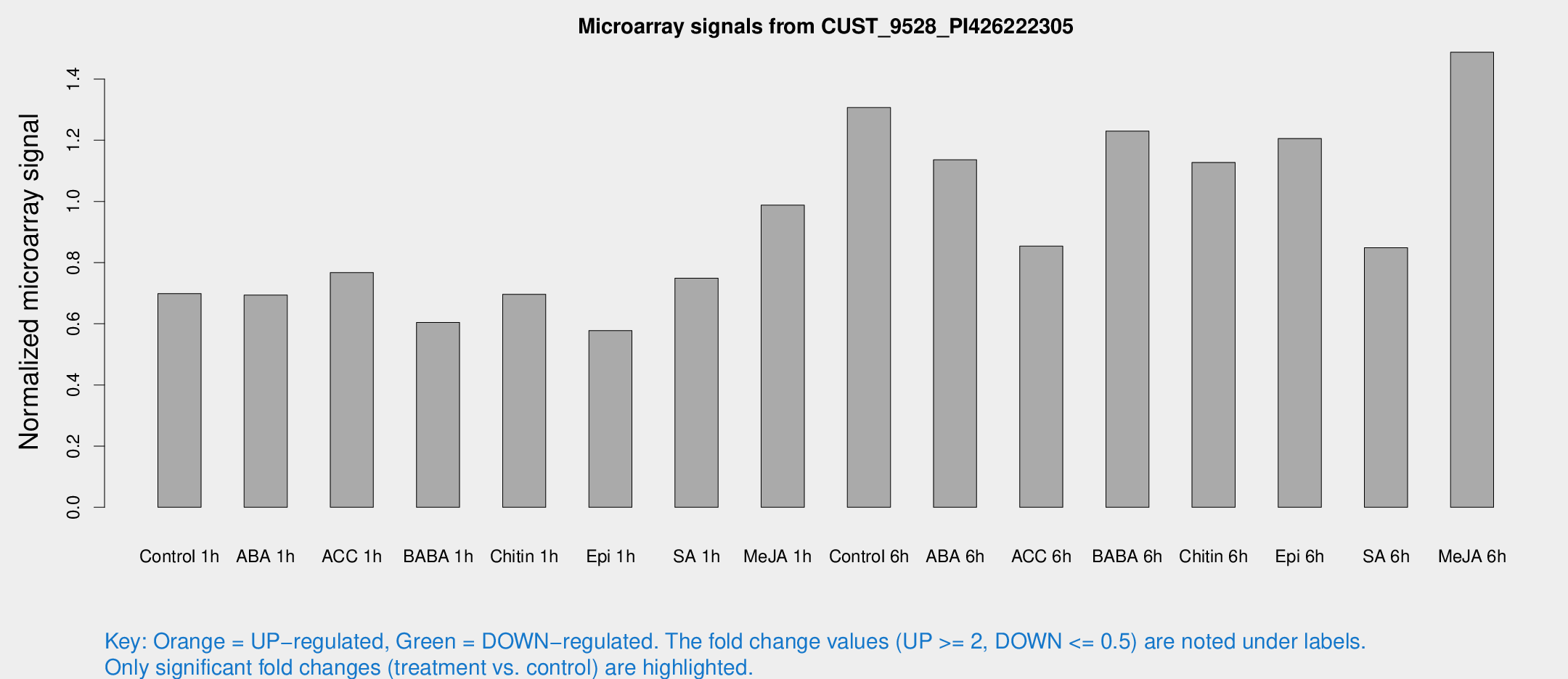
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 42.0623 | 14.3653 | 0.69892 | 0.271077 |
| ABA 1h | 37.6552 | 14.6019 | 0.693869 | 0.311793 |
| ACC 1h | 42.3678 | 8.33118 | 0.767411 | 0.177516 |
| BABA 1h | 36.4539 | 12.8216 | 0.604527 | 0.254912 |
| Chitin 1h | 34.686 | 8.85338 | 0.696438 | 0.134155 |
| Epi 1h | 30.6543 | 10.8406 | 0.577952 | 0.281939 |
| SA 1h | 40.7718 | 6.21834 | 0.749261 | 0.0796141 |
| Me-JA 1h | 44.1308 | 10.4459 | 0.987923 | 0.211522 |
| Control 6h | 68.5332 | 7.25543 | 1.30689 | 0.105333 |
| ABA 6h | 63.3969 | 7.87389 | 1.1363 | 0.112258 |
| ACC 6h | 52.716 | 10.5398 | 0.853743 | 0.202988 |
| BABA 6h | 72.0499 | 7.76748 | 1.23006 | 0.0999632 |
| Chitin 6h | 62.5296 | 5.64716 | 1.12752 | 0.0977369 |
| Epi 6h | 73.6373 | 16.0419 | 1.20581 | 0.315844 |
| SA 6h | 46.5859 | 12.3151 | 0.848847 | 0.156857 |
| Me-JA 6h | 80.383 | 18.0438 | 1.48784 | 0.2078 |
Source Transcript PGSC0003DMT400023617 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14205.1 | +3 | 0.0 | 885 | 478/822 (58%) | Phosphoinositide phosphatase family protein | chr3:4716008-4720524 REVERSE LENGTH=808 |
