Probe CUST_9465_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9465_PI426222305 | JHI_St_60k_v1 | DMT400023726 | CGAATTGGAGTATTGGAATTGTGTTTTATCAACTTCTATTCGTGTTGACGAATTTCAGTG |
All Microarray Probes Designed to Gene DMG400009178
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9465_PI426222305 | JHI_St_60k_v1 | DMT400023726 | CGAATTGGAGTATTGGAATTGTGTTTTATCAACTTCTATTCGTGTTGACGAATTTCAGTG |
| CUST_9178_PI426222305 | JHI_St_60k_v1 | DMT400023727 | ACTGTCAAGGCCGTTCGACGCAACTACTATGCAGTCAAGGAACTCATCAAAACTAGGAAA |
Microarray Signals from CUST_9465_PI426222305
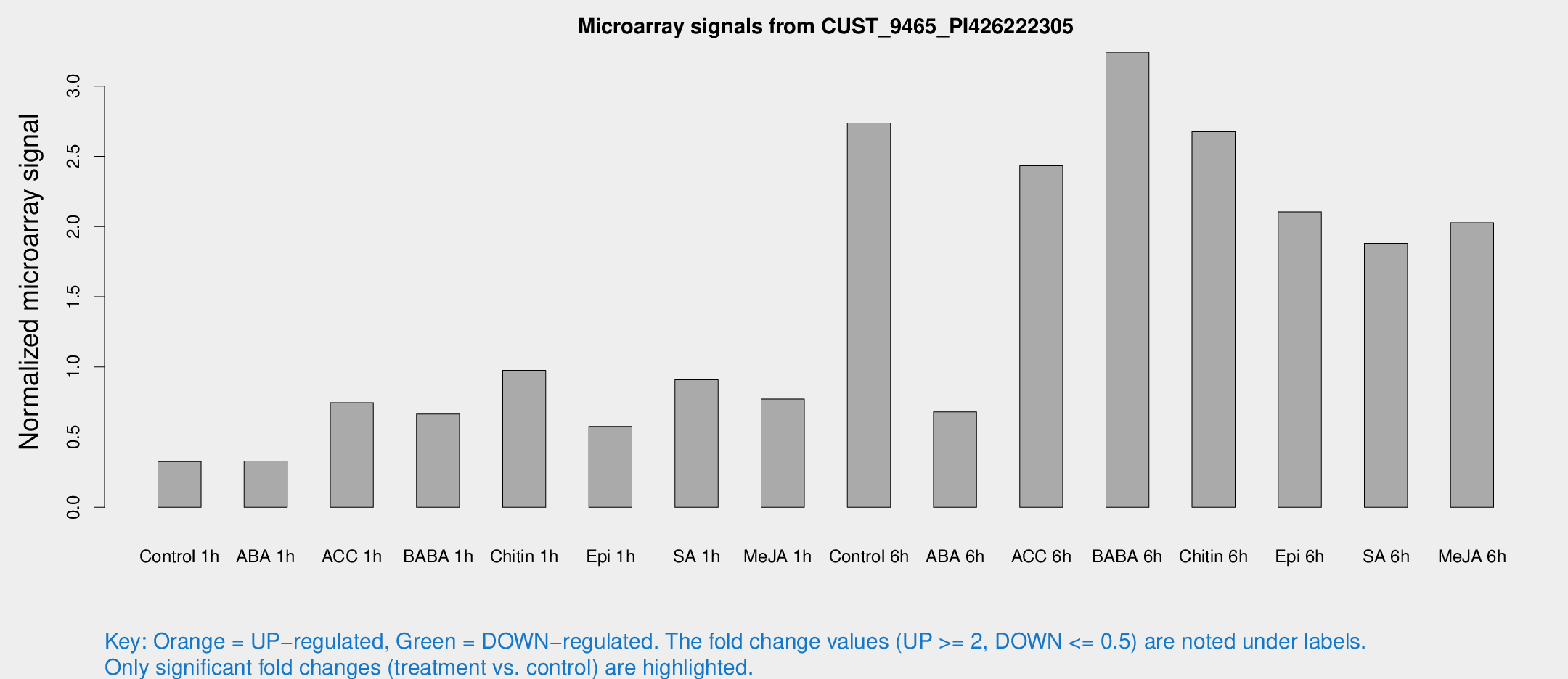
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2049.43 | 454.406 | 0.326129 | 0.0511701 |
| ABA 1h | 1824.9 | 366.239 | 0.329887 | 0.0668465 |
| ACC 1h | 4837.1 | 1123.71 | 0.745734 | 0.125362 |
| BABA 1h | 3833.88 | 221.622 | 0.663908 | 0.0488749 |
| Chitin 1h | 5342.34 | 729.717 | 0.975654 | 0.147812 |
| Epi 1h | 3097.04 | 575.882 | 0.576432 | 0.104447 |
| SA 1h | 5594.48 | 323.457 | 0.908769 | 0.0632652 |
| Me-JA 1h | 3782.1 | 218.935 | 0.772197 | 0.0944572 |
| Control 6h | 20011.9 | 8006.09 | 2.73846 | 1.14901 |
| ABA 6h | 4444.59 | 681.08 | 0.680137 | 0.117806 |
| ACC 6h | 17625 | 4190.78 | 2.43267 | 0.236591 |
| BABA 6h | 21885.3 | 1939.69 | 3.2418 | 0.187166 |
| Chitin 6h | 17181.2 | 1582.15 | 2.67534 | 0.245286 |
| Epi 6h | 15006.4 | 3160.06 | 2.10557 | 0.700115 |
| SA 6h | 11402.9 | 1858.28 | 1.87961 | 0.267074 |
| Me-JA 6h | 13913.9 | 4899.33 | 2.02719 | 0.659307 |
Source Transcript PGSC0003DMT400023726 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14310.1 | +2 | 1e-135 | 402 | 194/236 (82%) | pectin methylesterase 3 | chr3:4772214-4775095 REVERSE LENGTH=592 |
