Probe CUST_9375_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9375_PI426222305 | JHI_St_60k_v1 | DMT400023713 | ACCCCAAAAGAGTACATTTTCTATATGGATGTTCCTGGTTTATCCAAATCTGATATTCAG |
All Microarray Probes Designed to Gene DMG400009173
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9375_PI426222305 | JHI_St_60k_v1 | DMT400023713 | ACCCCAAAAGAGTACATTTTCTATATGGATGTTCCTGGTTTATCCAAATCTGATATTCAG |
Microarray Signals from CUST_9375_PI426222305
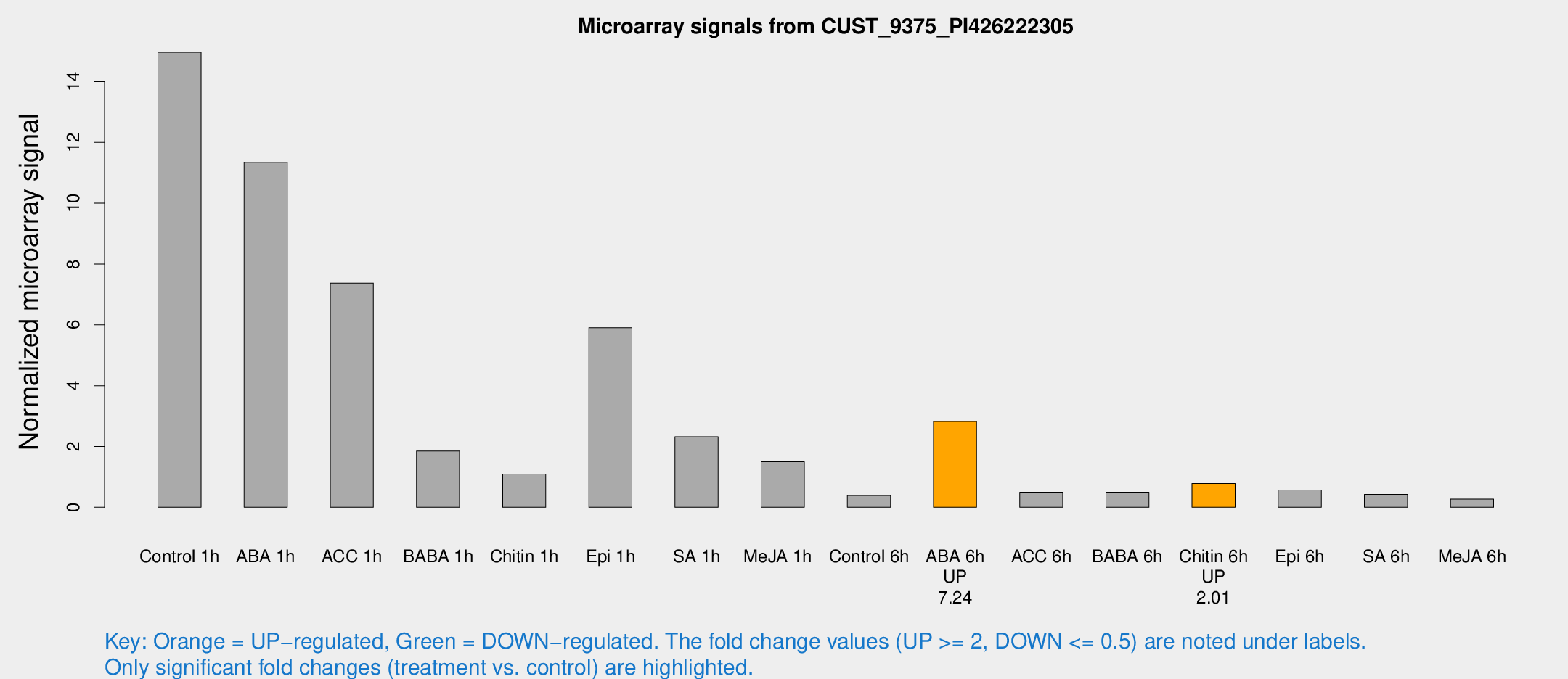
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1260.47 | 324.051 | 14.9647 | 3.6868 |
| ABA 1h | 1060.2 | 580.293 | 11.3426 | 7.90049 |
| ACC 1h | 808.589 | 397.882 | 7.37345 | 4.7075 |
| BABA 1h | 195.995 | 112.013 | 1.85102 | 1.10813 |
| Chitin 1h | 86.1488 | 29.3031 | 1.09277 | 0.279457 |
| Epi 1h | 475.762 | 169.257 | 5.90646 | 3.16359 |
| SA 1h | 200.001 | 44.6281 | 2.31885 | 0.509208 |
| Me-JA 1h | 98.3837 | 12.5551 | 1.49837 | 0.343035 |
| Control 6h | 32.5892 | 7.08806 | 0.389639 | 0.063141 |
| ABA 6h | 240.615 | 28.9449 | 2.82037 | 0.274427 |
| ACC 6h | 45.8875 | 6.63513 | 0.49512 | 0.0510032 |
| BABA 6h | 46.8744 | 12.3065 | 0.494681 | 0.144612 |
| Chitin 6h | 66.7389 | 7.78858 | 0.782663 | 0.111336 |
| Epi 6h | 51.6203 | 6.72275 | 0.570091 | 0.0662274 |
| SA 6h | 35.1094 | 7.55449 | 0.424258 | 0.0567444 |
| Me-JA 6h | 22.5577 | 5.56534 | 0.268272 | 0.0490185 |
Source Transcript PGSC0003DMT400023713 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G54050.1 | +1 | 3e-37 | 127 | 73/143 (51%) | HSP20-like chaperones superfamily protein | chr1:20179558-20180122 REVERSE LENGTH=155 |
