Probe CUST_9178_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9178_PI426222305 | JHI_St_60k_v1 | DMT400023727 | ACTGTCAAGGCCGTTCGACGCAACTACTATGCAGTCAAGGAACTCATCAAAACTAGGAAA |
All Microarray Probes Designed to Gene DMG400009178
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9465_PI426222305 | JHI_St_60k_v1 | DMT400023726 | CGAATTGGAGTATTGGAATTGTGTTTTATCAACTTCTATTCGTGTTGACGAATTTCAGTG |
| CUST_9178_PI426222305 | JHI_St_60k_v1 | DMT400023727 | ACTGTCAAGGCCGTTCGACGCAACTACTATGCAGTCAAGGAACTCATCAAAACTAGGAAA |
Microarray Signals from CUST_9178_PI426222305
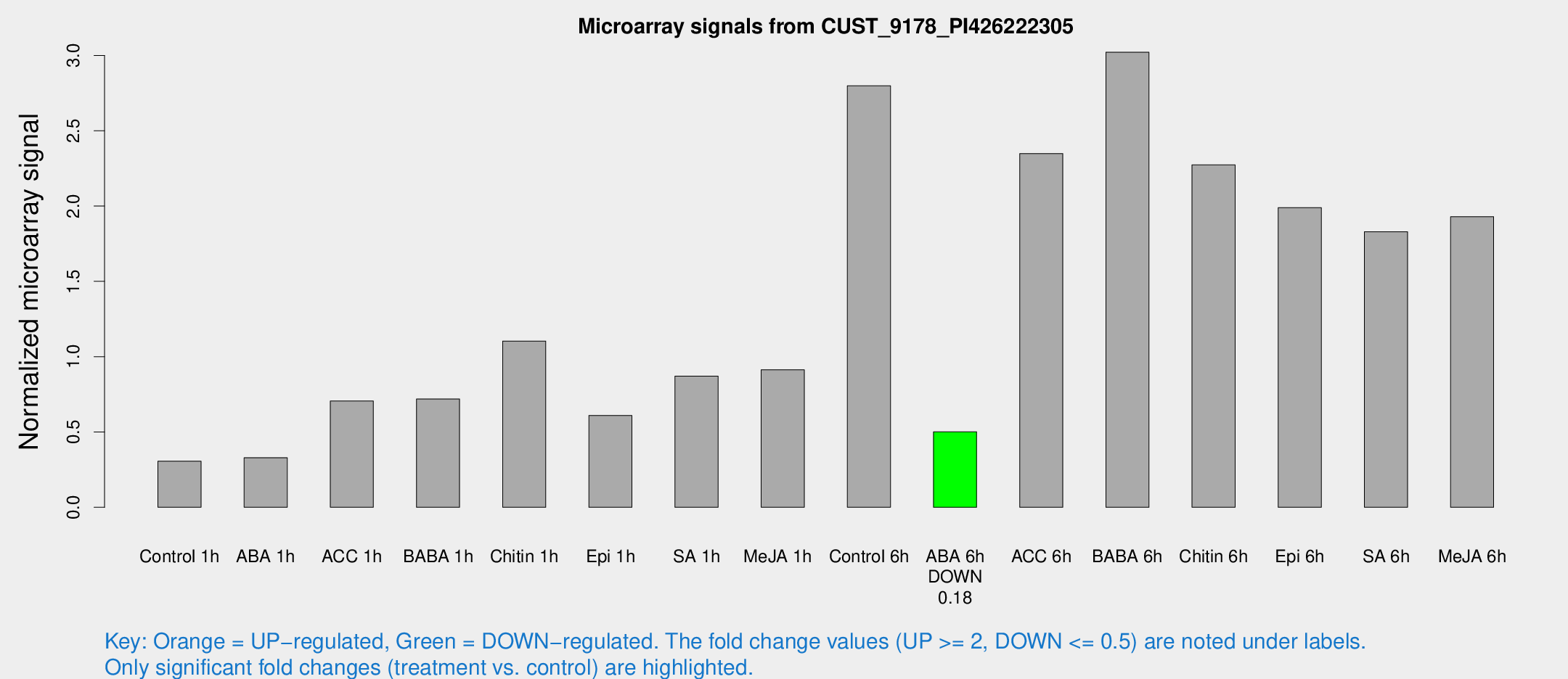
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 292.818 | 40.0108 | 0.306283 | 0.0229984 |
| ABA 1h | 278.48 | 36.5015 | 0.329439 | 0.0476877 |
| ACC 1h | 706.046 | 137.655 | 0.70589 | 0.0914597 |
| BABA 1h | 661.954 | 90.7659 | 0.719122 | 0.177512 |
| Chitin 1h | 1005.82 | 281.913 | 1.10377 | 0.380159 |
| Epi 1h | 508.743 | 79.5645 | 0.60994 | 0.097851 |
| SA 1h | 844.885 | 72.9663 | 0.870297 | 0.122794 |
| Me-JA 1h | 703.834 | 58.7207 | 0.913059 | 0.158913 |
| Control 6h | 3047.93 | 1102.58 | 2.79784 | 0.91859 |
| ABA 6h | 507.198 | 59.3653 | 0.501212 | 0.0718701 |
| ACC 6h | 2642.48 | 561.415 | 2.3485 | 0.214412 |
| BABA 6h | 3247.61 | 480.559 | 3.02108 | 0.357225 |
| Chitin 6h | 2272.49 | 131.425 | 2.27344 | 0.155064 |
| Epi 6h | 2405.55 | 741.382 | 1.98943 | 1.06626 |
| SA 6h | 2372.24 | 1125.59 | 1.82966 | 2.14069 |
| Me-JA 6h | 2102.84 | 741.714 | 1.9292 | 0.692356 |
Source Transcript PGSC0003DMT400023727 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14310.1 | +1 | 0.0 | 803 | 408/544 (75%) | pectin methylesterase 3 | chr3:4772214-4775095 REVERSE LENGTH=592 |
