Probe CUST_9165_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9165_PI426222305 | JHI_St_60k_v1 | DMT400023618 | GGACTGTCGCTTTAGTATATGGATTCTTTAAACAGAACTTCTATATGGTTGGAAGAGATA |
All Microarray Probes Designed to Gene DMG400009144
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9165_PI426222305 | JHI_St_60k_v1 | DMT400023618 | GGACTGTCGCTTTAGTATATGGATTCTTTAAACAGAACTTCTATATGGTTGGAAGAGATA |
| CUST_9528_PI426222305 | JHI_St_60k_v1 | DMT400023617 | GCTATCCAAAATGCCTTATTCTAAGAATGAGAACAGAACTTCTATATGGTTGGAAGAGAT |
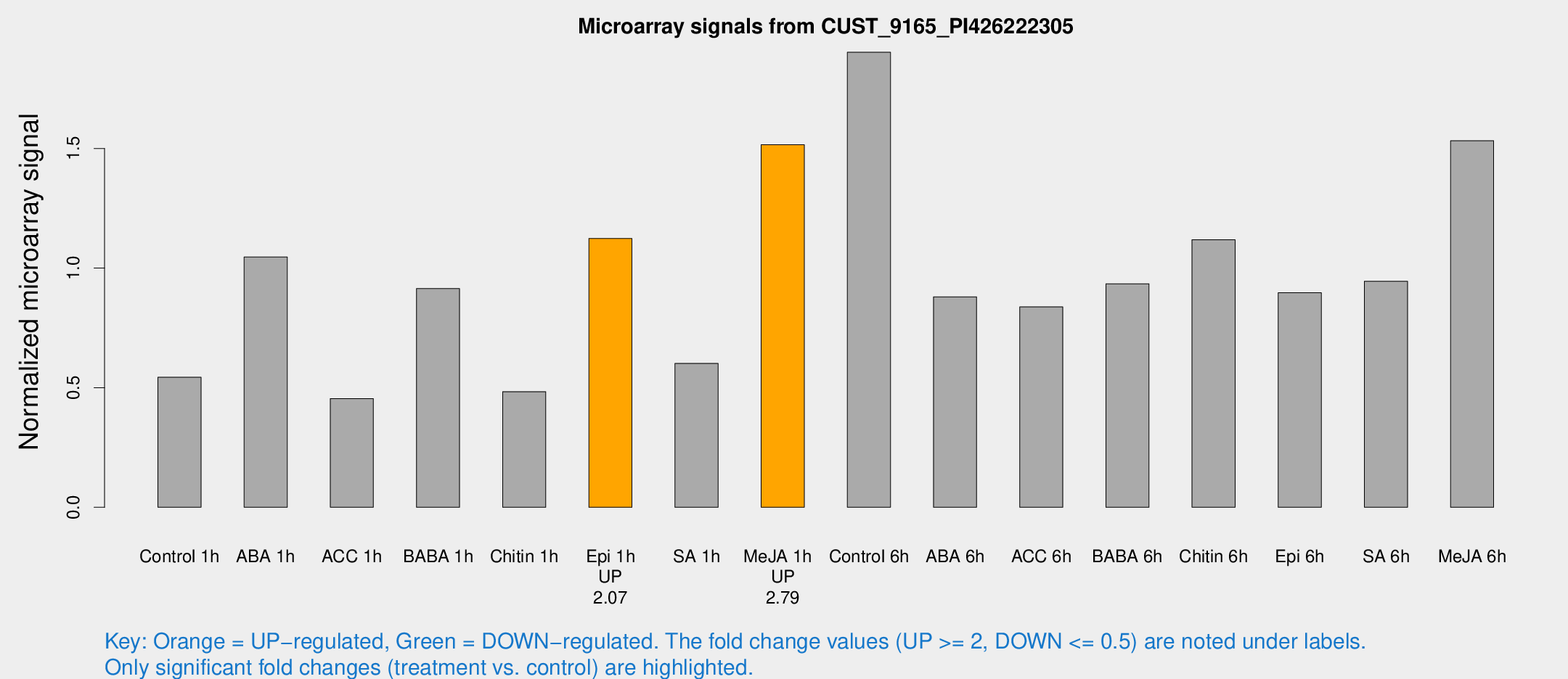
Microarray Signals from CUST_9165_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.98021 | 3.10887 | 0.543607 | 0.257103 |
| ABA 1h | 12.4319 | 3.3107 | 1.04627 | 0.313236 |
| ACC 1h | 5.81011 | 3.37309 | 0.45467 | 0.263639 |
| BABA 1h | 12.1475 | 3.39318 | 0.914444 | 0.327029 |
| Chitin 1h | 5.30048 | 3.08223 | 0.483473 | 0.273841 |
| Epi 1h | 12.0982 | 3.14343 | 1.12386 | 0.292043 |
| SA 1h | 8.11994 | 3.09689 | 0.60165 | 0.257664 |
| Me-JA 1h | 16.1104 | 3.27977 | 1.51562 | 0.335858 |
| Control 6h | 29.0538 | 10.8368 | 1.90256 | 0.878386 |
| ABA 6h | 13.3225 | 4.16059 | 0.880129 | 0.336757 |
| ACC 6h | 13.6346 | 4.16555 | 0.837999 | 0.304768 |
| BABA 6h | 14.9771 | 4.6503 | 0.934626 | 0.388252 |
| Chitin 6h | 15.7653 | 3.76361 | 1.11873 | 0.295046 |
| Epi 6h | 13.4261 | 3.7127 | 0.897434 | 0.330746 |
| SA 6h | 11.6826 | 3.36729 | 0.945286 | 0.276346 |
| Me-JA 6h | 26.0062 | 10.5063 | 1.53256 | 1.14335 |
Source Transcript PGSC0003DMT400023618 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14205.1 | +3 | 3e-126 | 415 | 231/395 (58%) | Phosphoinositide phosphatase family protein | chr3:4716008-4720524 REVERSE LENGTH=808 |
