Probe CUST_9115_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9115_PI426222305 | JHI_St_60k_v1 | DMT400095180 | CAGTTGTTGAAGGAACATGAAGTTGGAGATTCACTTAAAGGGTTGATTAATTTTCTTCAC |
All Microarray Probes Designed to Gene DMG400044751
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9115_PI426222305 | JHI_St_60k_v1 | DMT400095180 | CAGTTGTTGAAGGAACATGAAGTTGGAGATTCACTTAAAGGGTTGATTAATTTTCTTCAC |
Microarray Signals from CUST_9115_PI426222305
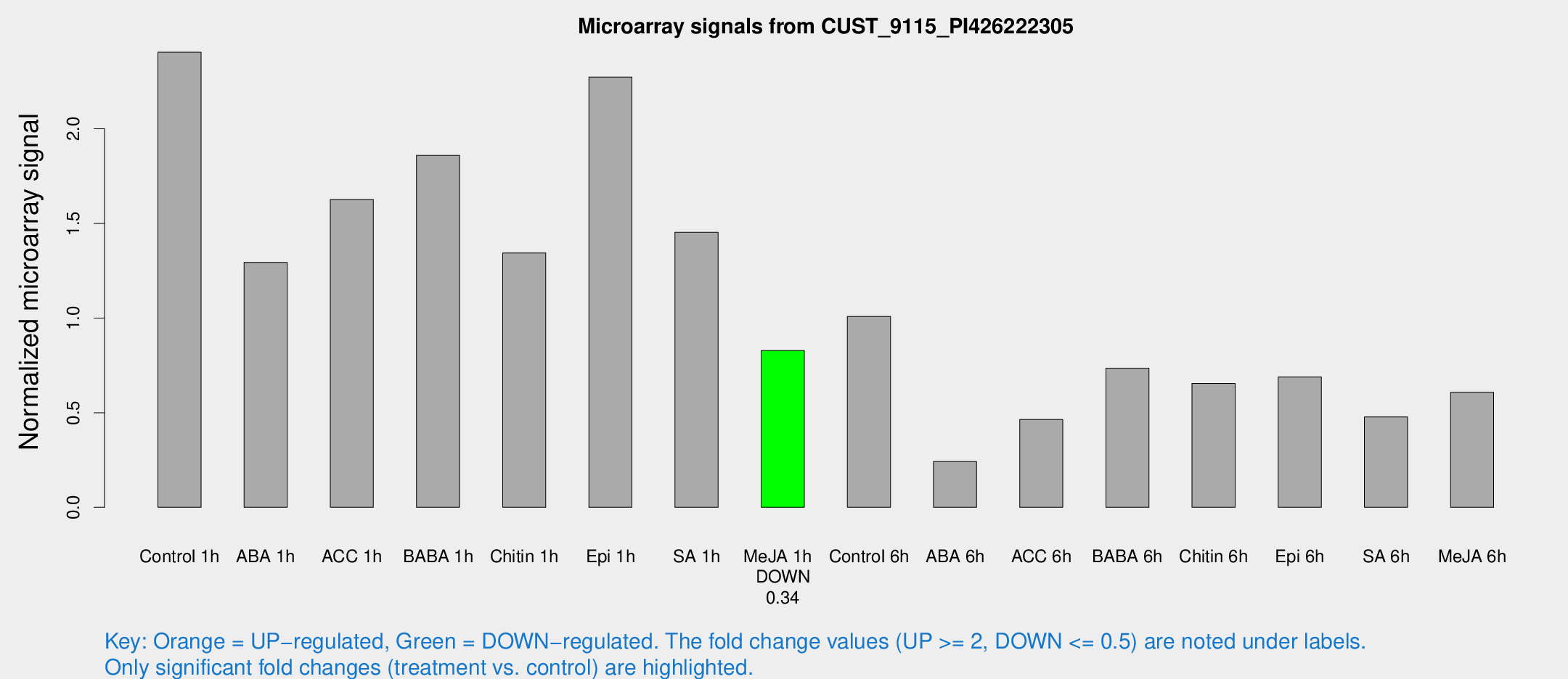
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 174.025 | 19.8923 | 2.40472 | 0.158639 |
| ABA 1h | 82.8512 | 8.78054 | 1.29445 | 0.0886531 |
| ACC 1h | 178.459 | 88.2566 | 1.62618 | 1.46769 |
| BABA 1h | 129.079 | 10.8546 | 1.85986 | 0.117578 |
| Chitin 1h | 86.6576 | 5.88603 | 1.34435 | 0.0912092 |
| Epi 1h | 140.872 | 8.70153 | 2.27347 | 0.140276 |
| SA 1h | 107.666 | 10.2708 | 1.45276 | 0.204377 |
| Me-JA 1h | 50.6783 | 11.4364 | 0.828629 | 0.129514 |
| Control 6h | 84.0071 | 28.3806 | 1.00842 | 0.360933 |
| ABA 6h | 21.0812 | 6.46254 | 0.241831 | 0.0950754 |
| ACC 6h | 39.0492 | 6.40637 | 0.464128 | 0.0519721 |
| BABA 6h | 65.9579 | 19.773 | 0.73552 | 0.258446 |
| Chitin 6h | 50.2211 | 4.60925 | 0.65442 | 0.0594742 |
| Epi 6h | 56.5962 | 7.66317 | 0.68799 | 0.148626 |
| SA 6h | 40.8622 | 17.1686 | 0.477691 | 0.255475 |
| Me-JA 6h | 47.2665 | 13.1541 | 0.607969 | 0.139478 |
Source Transcript PGSC0003DMT400095180 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G50940.1 | +1 | 2e-21 | 88 | 41/69 (59%) | P-loop containing nucleoside triphosphate hydrolases superfamily protein | chr3:18934086-18935528 FORWARD LENGTH=451 |
