Probe CUST_8997_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8997_PI426222305 | JHI_St_60k_v1 | DMT400013020 | GATAGACCCTTGTTGTCCTATTCTTGTTGATTTTTCATTTTTAGGCATGTATTCGACTAC |
All Microarray Probes Designed to Gene DMG400005073
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8997_PI426222305 | JHI_St_60k_v1 | DMT400013020 | GATAGACCCTTGTTGTCCTATTCTTGTTGATTTTTCATTTTTAGGCATGTATTCGACTAC |
Microarray Signals from CUST_8997_PI426222305
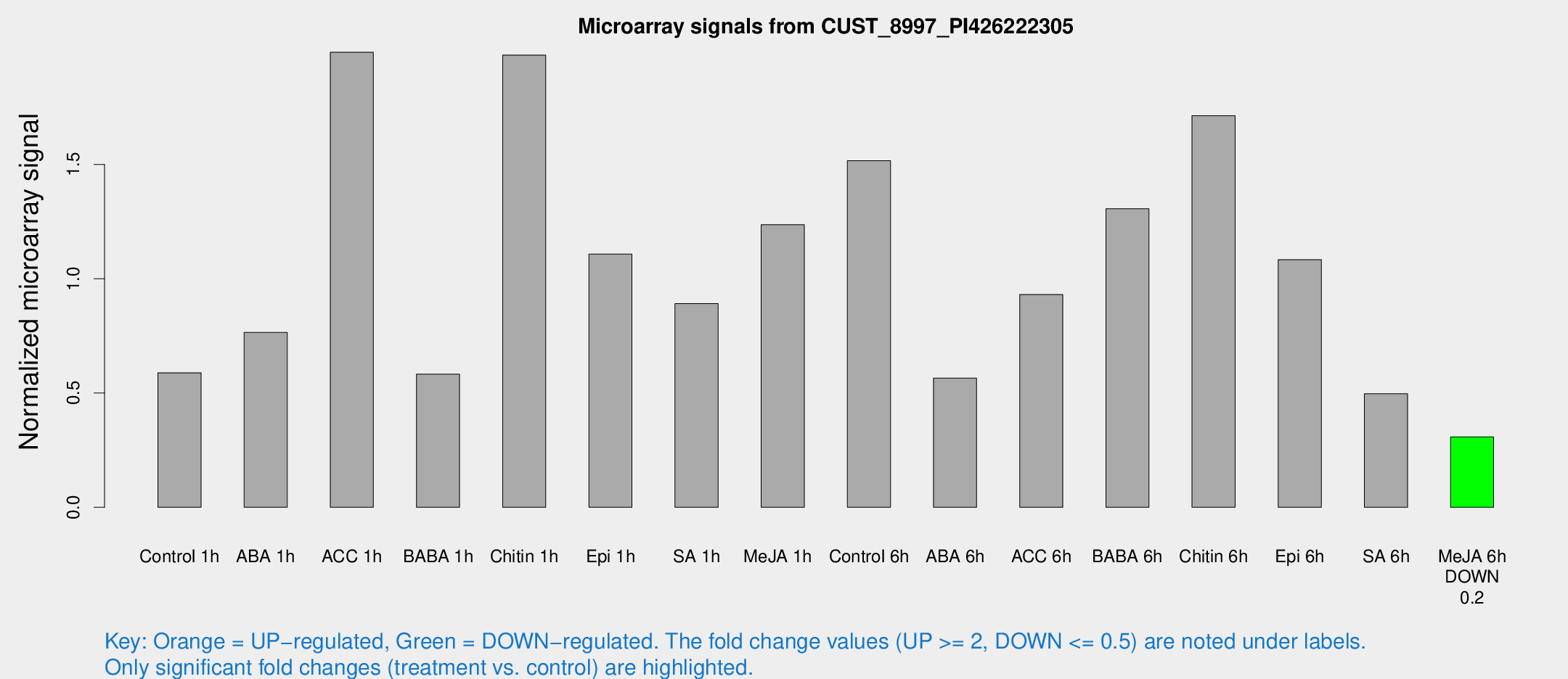
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 14.8314 | 5.21821 | 0.588482 | 0.229694 |
| ABA 1h | 20.9011 | 11.9797 | 0.765181 | 0.702837 |
| ACC 1h | 48.0267 | 13.0594 | 1.99135 | 0.434401 |
| BABA 1h | 13.5355 | 4.06587 | 0.582698 | 0.266666 |
| Chitin 1h | 52.6385 | 26.0874 | 1.9788 | 1.68772 |
| Epi 1h | 23.786 | 8.26206 | 1.10786 | 0.435507 |
| SA 1h | 25.5929 | 9.61588 | 0.8913 | 0.728079 |
| Me-JA 1h | 26.9012 | 9.46991 | 1.2363 | 0.890665 |
| Control 6h | 39.0267 | 15.2656 | 1.51668 | 0.510648 |
| ABA 6h | 16.7495 | 8.446 | 0.565061 | 0.367624 |
| ACC 6h | 24.2076 | 4.78035 | 0.930852 | 0.183762 |
| BABA 6h | 32.4631 | 4.75141 | 1.30657 | 0.199736 |
| Chitin 6h | 44.1156 | 11.9023 | 1.7138 | 0.565067 |
| Epi 6h | 33.8139 | 12.7684 | 1.08384 | 0.858732 |
| SA 6h | 12.9716 | 5.81744 | 0.497003 | 0.226016 |
| Me-JA 6h | 6.75751 | 3.92075 | 0.308339 | 0.178556 |
Source Transcript PGSC0003DMT400013020 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G61760.1 | +1 | 1e-67 | 216 | 110/226 (49%) | Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family | chr1:22807440-22808114 REVERSE LENGTH=224 |
