Probe CUST_8549_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8549_PI426222305 | JHI_St_60k_v1 | DMT400027659 | GAAACCGTAATATCGCTATAGTAATGAATAGGTAGTAGCTGAAAGTTGCAATCATTCAGT |
All Microarray Probes Designed to Gene DMG400010664
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8536_PI426222305 | JHI_St_60k_v1 | DMT400027660 | GAAACCGTAATATCGCTATAGTAATGAATAGGTAGTAGCTGAAAGTTGCAATCATTCAGT |
| CUST_8549_PI426222305 | JHI_St_60k_v1 | DMT400027659 | GAAACCGTAATATCGCTATAGTAATGAATAGGTAGTAGCTGAAAGTTGCAATCATTCAGT |
Microarray Signals from CUST_8549_PI426222305
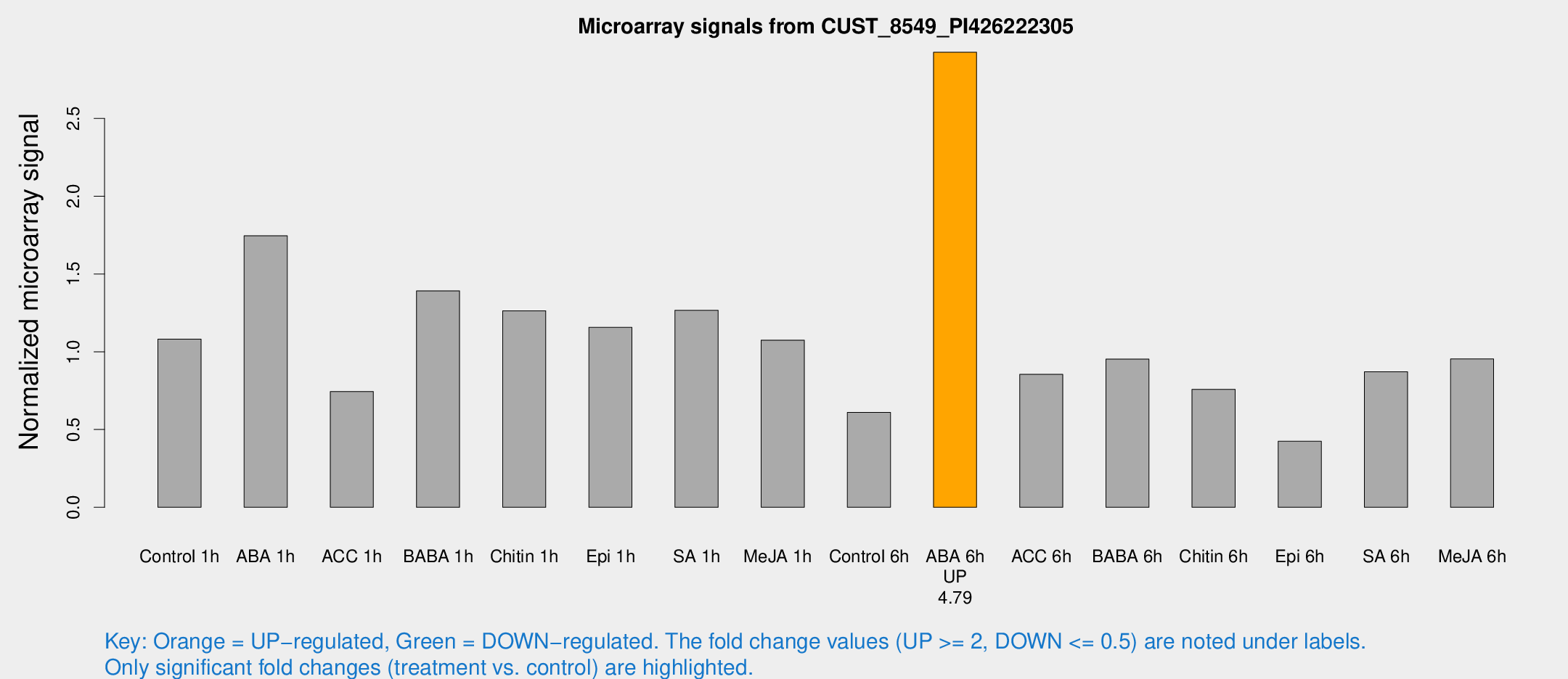
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1156.44 | 102.5 | 1.08067 | 0.0624866 |
| ABA 1h | 1763.58 | 437.808 | 1.7458 | 0.441346 |
| ACC 1h | 922.165 | 290.858 | 0.744463 | 0.246041 |
| BABA 1h | 1551.13 | 394.659 | 1.39161 | 0.295478 |
| Chitin 1h | 1311.11 | 388.997 | 1.26338 | 0.316324 |
| Epi 1h | 1093.96 | 173.54 | 1.15704 | 0.186908 |
| SA 1h | 1441.32 | 312.284 | 1.26646 | 0.191247 |
| Me-JA 1h | 964.384 | 186.806 | 1.0748 | 0.143488 |
| Control 6h | 706.018 | 198.443 | 0.610477 | 0.14195 |
| ABA 6h | 3330.49 | 314.745 | 2.9258 | 0.219455 |
| ACC 6h | 1045.74 | 76.2306 | 0.854584 | 0.05785 |
| BABA 6h | 1169.7 | 220.079 | 0.952455 | 0.139856 |
| Chitin 6h | 859.601 | 49.9377 | 0.75874 | 0.0439585 |
| Epi 6h | 516.984 | 70.3242 | 0.424642 | 0.0357631 |
| SA 6h | 963.44 | 199.522 | 0.871723 | 0.104089 |
| Me-JA 6h | 1035.35 | 163.764 | 0.953744 | 0.0778463 |
Source Transcript PGSC0003DMT400027659 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G18670.1 | +3 | 0.0 | 579 | 297/524 (57%) | beta-amylase 3 | chr5:6226138-6227999 FORWARD LENGTH=536 |
