Probe CUST_7815_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_7815_PI426222305 | JHI_St_60k_v1 | DMT400025890 | CTTTCACAGCAGAGACTACCTGATATTGCAACATTATTCAGCCAGTTTCTGTTTTATTTT |
All Microarray Probes Designed to Gene DMG401009997
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_7815_PI426222305 | JHI_St_60k_v1 | DMT400025890 | CTTTCACAGCAGAGACTACCTGATATTGCAACATTATTCAGCCAGTTTCTGTTTTATTTT |
Microarray Signals from CUST_7815_PI426222305
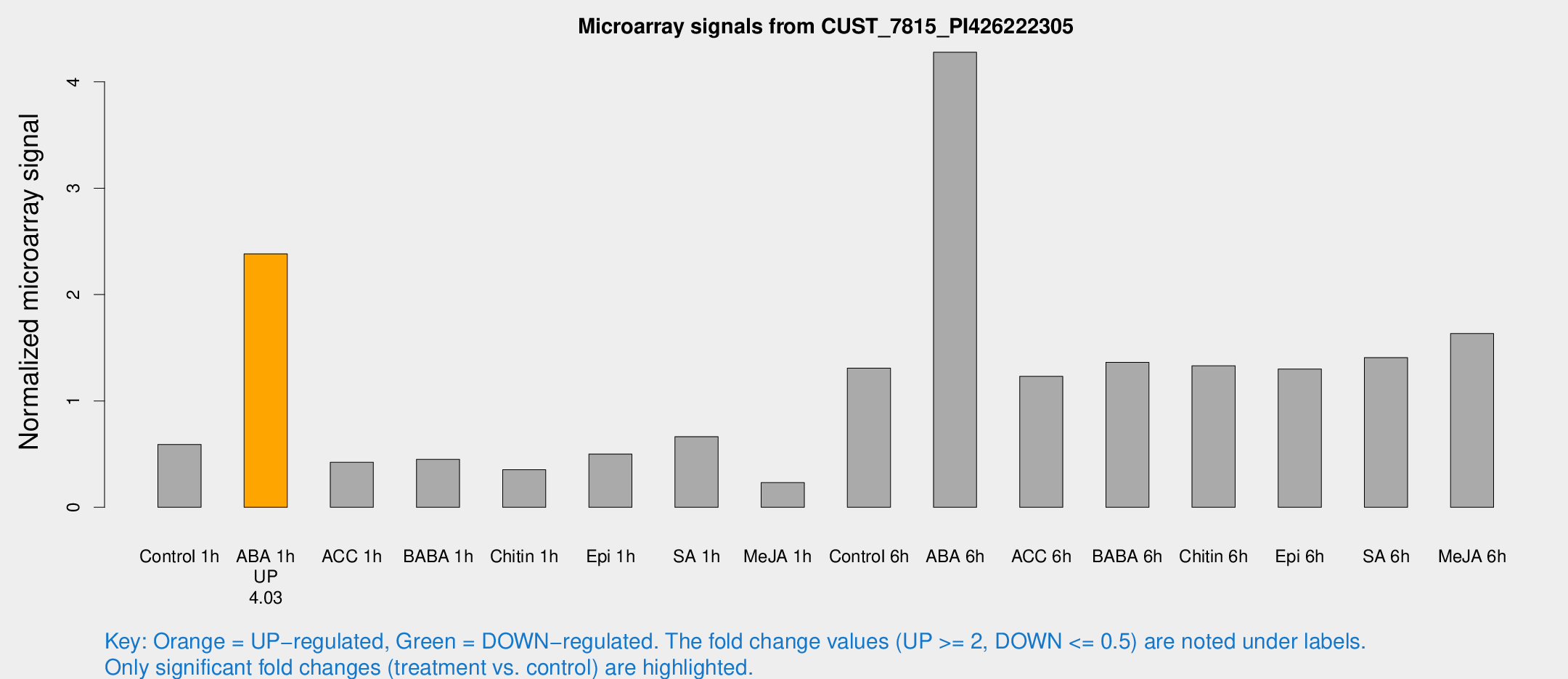
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 32.5219 | 6.5736 | 0.591325 | 0.104233 |
| ABA 1h | 111.422 | 7.29546 | 2.3825 | 0.156113 |
| ACC 1h | 29.512 | 11.1089 | 0.42317 | 0.25136 |
| BABA 1h | 23.2116 | 3.97508 | 0.451441 | 0.0789641 |
| Chitin 1h | 16.9768 | 3.72304 | 0.353644 | 0.0794452 |
| Epi 1h | 25.8303 | 7.5814 | 0.500553 | 0.198601 |
| SA 1h | 36.1803 | 3.91926 | 0.664218 | 0.0727015 |
| Me-JA 1h | 13.3546 | 7.38141 | 0.232169 | 0.119491 |
| Control 6h | 74.4459 | 20.9058 | 1.30818 | 0.363128 |
| ABA 6h | 245.399 | 34.2202 | 4.27874 | 0.508032 |
| ACC 6h | 74.612 | 6.2586 | 1.23169 | 0.167882 |
| BABA 6h | 80.971 | 6.24119 | 1.36333 | 0.124191 |
| Chitin 6h | 76.5205 | 11.1645 | 1.33044 | 0.197958 |
| Epi 6h | 79.3308 | 11.8204 | 1.3006 | 0.108276 |
| SA 6h | 76.2819 | 15.3892 | 1.40685 | 0.250957 |
| Me-JA 6h | 89.6979 | 19.0181 | 1.63326 | 0.208355 |
Source Transcript PGSC0003DMT400025890 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G54850.1 | +2 | 2e-15 | 76 | 43/118 (36%) | HSP20-like chaperones superfamily protein | chr1:20454440-20455994 FORWARD LENGTH=206 |
