Probe CUST_7680_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_7680_PI426222305 | JHI_St_60k_v1 | DMT400009594 | GCATATAGCGGGAGCTTTAATGTACCAGATTGTCCTTTCTTCTTCTTTTTATTTAAGTTG |
All Microarray Probes Designed to Gene DMG400003743
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_7680_PI426222305 | JHI_St_60k_v1 | DMT400009594 | GCATATAGCGGGAGCTTTAATGTACCAGATTGTCCTTTCTTCTTCTTTTTATTTAAGTTG |
Microarray Signals from CUST_7680_PI426222305
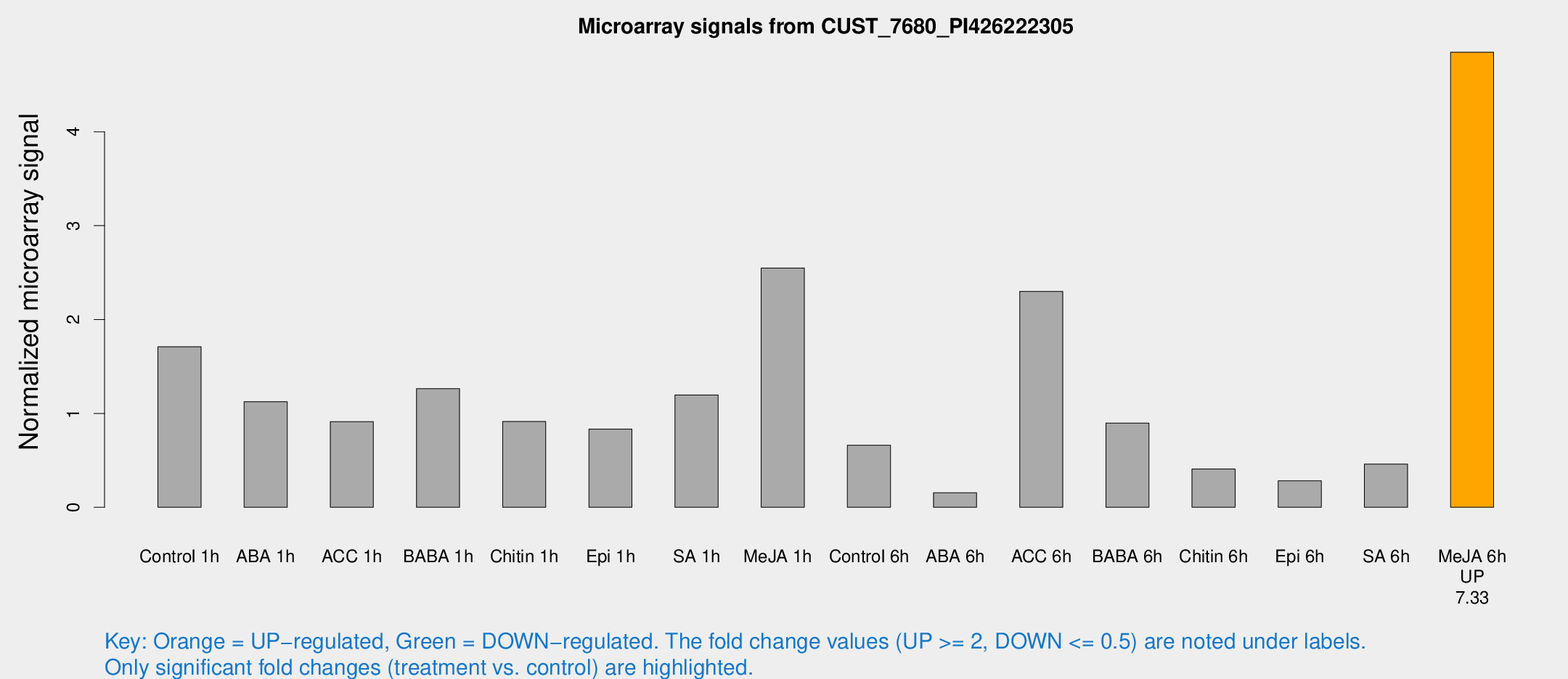
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 107.234 | 34.1763 | 1.70986 | 0.479329 |
| ABA 1h | 62.2827 | 18.6949 | 1.12565 | 0.362686 |
| ACC 1h | 65.4786 | 31.5714 | 0.912961 | 0.452577 |
| BABA 1h | 73.1611 | 16.7456 | 1.2633 | 0.208132 |
| Chitin 1h | 49.8657 | 12.7791 | 0.913742 | 0.228761 |
| Epi 1h | 53.4641 | 20.6827 | 0.832669 | 0.644899 |
| SA 1h | 71.823 | 13.0593 | 1.19614 | 0.193571 |
| Me-JA 1h | 119.573 | 16.9009 | 2.54813 | 0.164053 |
| Control 6h | 47.9403 | 24.7746 | 0.660907 | 0.344587 |
| ABA 6h | 10.3041 | 3.5905 | 0.15532 | 0.0657236 |
| ACC 6h | 154.934 | 32.793 | 2.29944 | 0.205919 |
| BABA 6h | 64.9887 | 25.416 | 0.897038 | 0.325798 |
| Chitin 6h | 24.853 | 4.03515 | 0.407469 | 0.0694744 |
| Epi 6h | 21.6936 | 8.63869 | 0.282506 | 0.131358 |
| SA 6h | 32.5819 | 13.2191 | 0.459874 | 0.222944 |
| Me-JA 6h | 292.638 | 71.1309 | 4.84702 | 1.86352 |
Source Transcript PGSC0003DMT400009594 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G19640.1 | +1 | 9e-125 | 372 | 201/388 (52%) | jasmonic acid carboxyl methyltransferase | chr1:6789166-6791708 REVERSE LENGTH=389 |
