Probe CUST_7515_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_7515_PI426222305 | JHI_St_60k_v1 | DMT400025631 | GTTTCATCTCGTGGGATTGTACTAGGTATGTTGTTGTTGTAGTCATAATTGTATTCTATG |
All Microarray Probes Designed to Gene DMG400009899
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_7540_PI426222305 | JHI_St_60k_v1 | DMT400025630 | GAGAACATGACTTCAGTTACTTCTCTTGATATCTCAAGGAATTCTCTTGTAGGCAAATAA |
| CUST_7515_PI426222305 | JHI_St_60k_v1 | DMT400025631 | GTTTCATCTCGTGGGATTGTACTAGGTATGTTGTTGTTGTAGTCATAATTGTATTCTATG |
Microarray Signals from CUST_7515_PI426222305
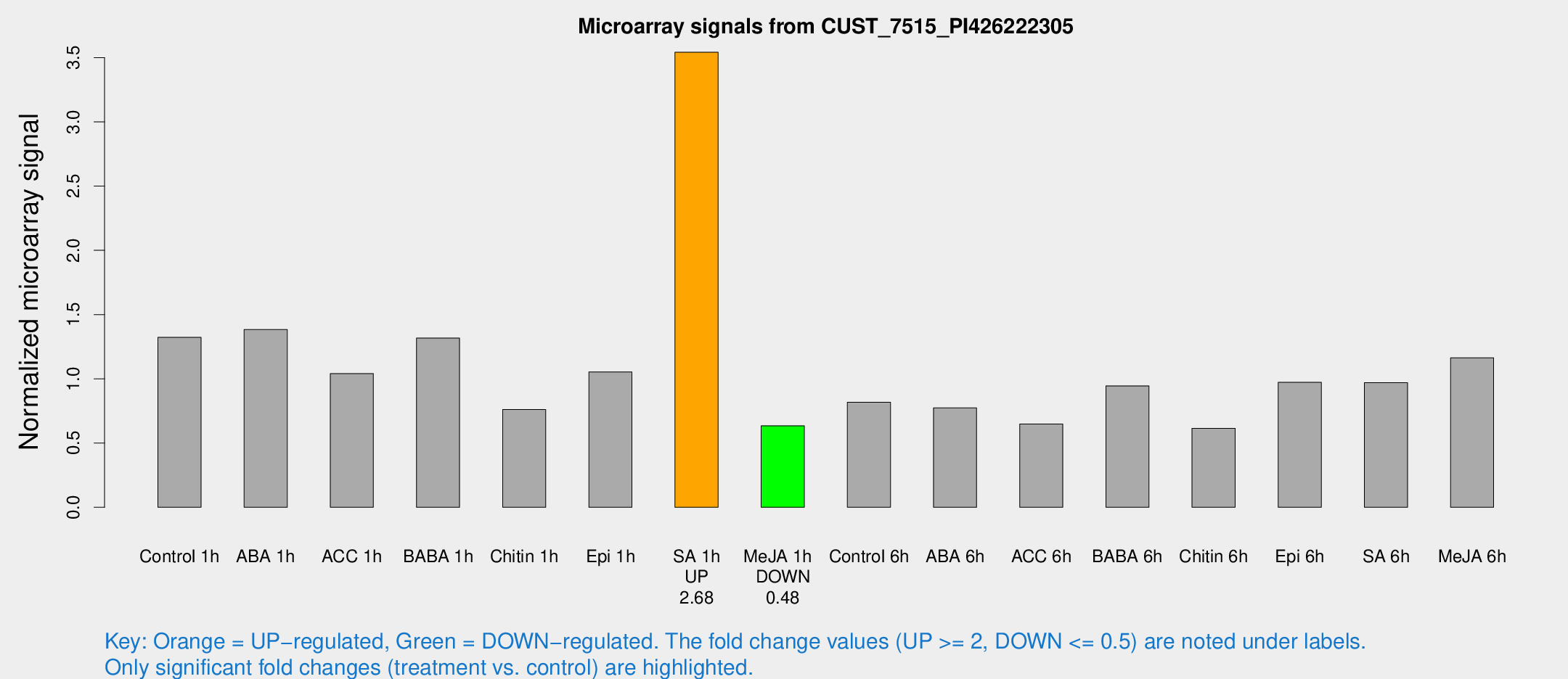
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 426.298 | 58.1915 | 1.32258 | 0.1084 |
| ABA 1h | 397.277 | 60.0285 | 1.38368 | 0.127536 |
| ACC 1h | 429.586 | 170.281 | 1.04074 | 0.573787 |
| BABA 1h | 438.126 | 115.047 | 1.31734 | 0.278155 |
| Chitin 1h | 249.701 | 92.1342 | 0.761688 | 0.282976 |
| Epi 1h | 296.001 | 45.4066 | 1.05365 | 0.164392 |
| SA 1h | 1178.56 | 179.904 | 3.54199 | 0.314738 |
| Me-JA 1h | 172.56 | 40.5209 | 0.63463 | 0.093657 |
| Control 6h | 269.137 | 47.552 | 0.81715 | 0.0877265 |
| ABA 6h | 298.463 | 108.325 | 0.773664 | 0.263269 |
| ACC 6h | 239.793 | 31.6605 | 0.648488 | 0.180256 |
| BABA 6h | 339.692 | 38.9482 | 0.945198 | 0.108379 |
| Chitin 6h | 207.961 | 12.6681 | 0.615088 | 0.0458782 |
| Epi 6h | 356.674 | 54.5896 | 0.972783 | 0.0906505 |
| SA 6h | 387.941 | 157.444 | 0.969565 | 0.479162 |
| Me-JA 6h | 415.121 | 149.449 | 1.16368 | 0.325608 |
Source Transcript PGSC0003DMT400025631 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G34930.1 | +1 | 2e-114 | 377 | 306/889 (34%) | disease resistance family protein / LRR family protein | chr2:14737169-14739886 REVERSE LENGTH=905 |
