Probe CUST_7381_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_7381_PI426222305 | JHI_St_60k_v1 | DMT400025690 | CCCTTTAGCTATATGTAATTAATTCAGGTTGCTTTATTCTGAAACCTGCATGTTGCTTGT |
All Microarray Probes Designed to Gene DMG400009920
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_7381_PI426222305 | JHI_St_60k_v1 | DMT400025690 | CCCTTTAGCTATATGTAATTAATTCAGGTTGCTTTATTCTGAAACCTGCATGTTGCTTGT |
Microarray Signals from CUST_7381_PI426222305
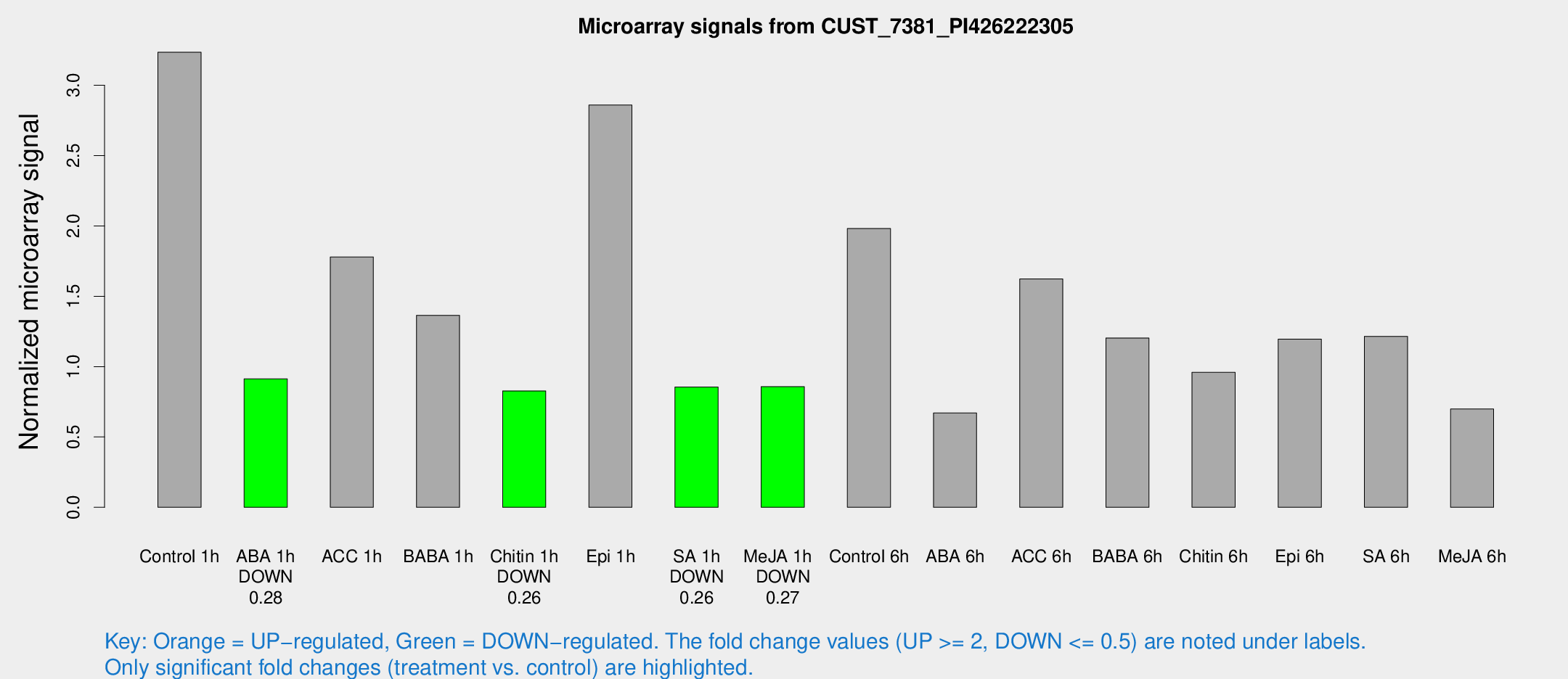
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 27.7882 | 4.88677 | 3.23511 | 0.698148 |
| ABA 1h | 6.94122 | 3.183 | 0.912667 | 0.44903 |
| ACC 1h | 16.155 | 4.1463 | 1.77944 | 0.600187 |
| BABA 1h | 12.0884 | 3.84567 | 1.36505 | 0.513267 |
| Chitin 1h | 6.17883 | 3.26203 | 0.827382 | 0.440082 |
| Epi 1h | 20.8368 | 3.42307 | 2.85972 | 0.497246 |
| SA 1h | 7.57737 | 3.31718 | 0.854808 | 0.407249 |
| Me-JA 1h | 5.81345 | 3.3786 | 0.857698 | 0.497241 |
| Control 6h | 23.7235 | 13.4464 | 1.98155 | 1.40866 |
| ABA 6h | 5.91295 | 3.42833 | 0.670683 | 0.388365 |
| ACC 6h | 16.1993 | 4.27641 | 1.62323 | 0.436541 |
| BABA 6h | 15.6185 | 9.2172 | 1.20338 | 0.843515 |
| Chitin 6h | 9.12007 | 3.72032 | 0.959518 | 0.459468 |
| Epi 6h | 13.4583 | 6.08139 | 1.19515 | 0.700602 |
| SA 6h | 10.3197 | 3.57416 | 1.21538 | 0.459589 |
| Me-JA 6h | 5.76858 | 3.34504 | 0.69874 | 0.404779 |
Source Transcript PGSC0003DMT400025690 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G76420.1 | +1 | 3e-95 | 293 | 159/269 (59%) | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein | chr1:28672029-28673835 REVERSE LENGTH=334 |
