Probe CUST_6707_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6707_PI426222305 | JHI_St_60k_v1 | DMT400014695 | TGTTGTTTCTGAGCAAATCAAAACTCGCCTTGAAGTGTCTCAACAAGAAACAAATACAAG |
All Microarray Probes Designed to Gene DMG400005734
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6707_PI426222305 | JHI_St_60k_v1 | DMT400014695 | TGTTGTTTCTGAGCAAATCAAAACTCGCCTTGAAGTGTCTCAACAAGAAACAAATACAAG |
Microarray Signals from CUST_6707_PI426222305
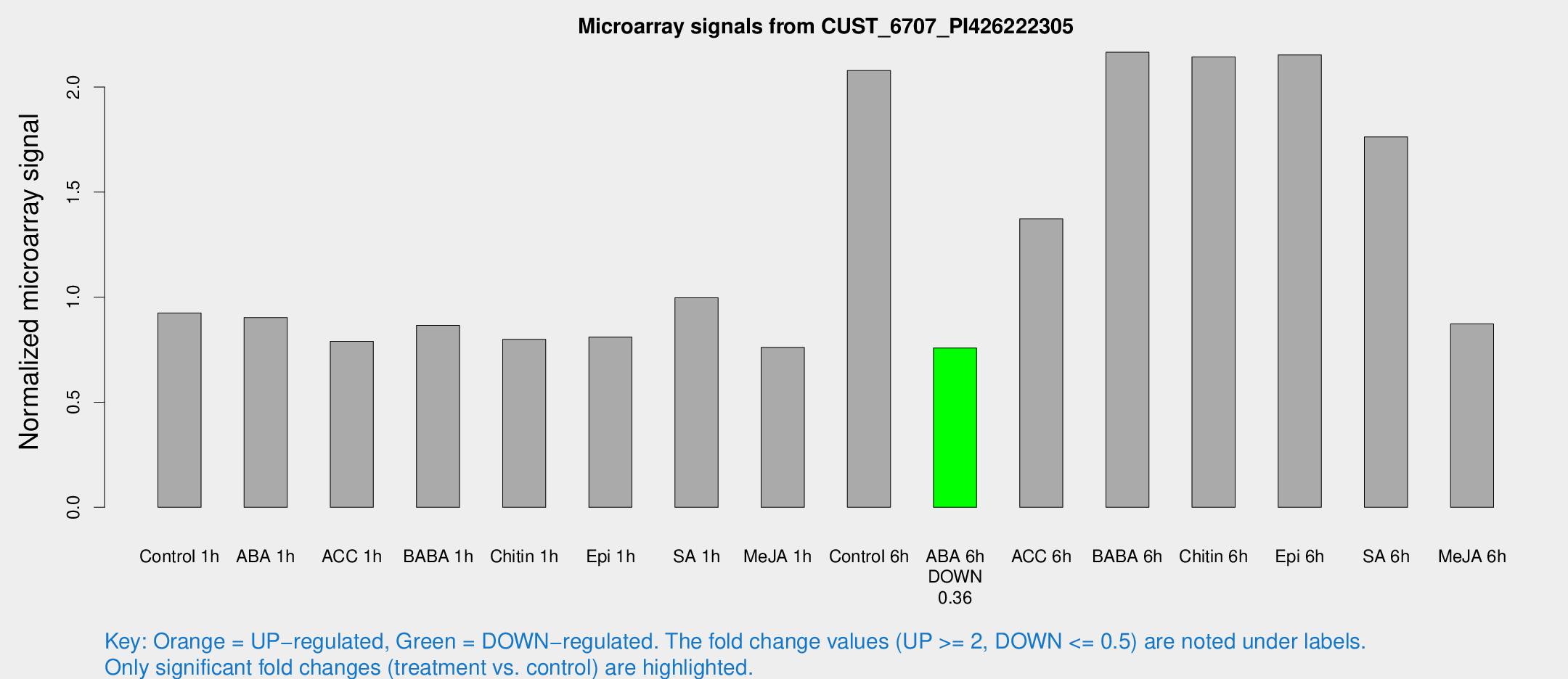
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2085.57 | 319.622 | 0.924559 | 0.0947022 |
| ABA 1h | 1793.07 | 236.961 | 0.902907 | 0.0930799 |
| ACC 1h | 1834.42 | 275.289 | 0.789606 | 0.0653802 |
| BABA 1h | 1926.79 | 377.84 | 0.865879 | 0.101965 |
| Chitin 1h | 1590.79 | 92.172 | 0.799241 | 0.104061 |
| Epi 1h | 1569.08 | 178.519 | 0.809916 | 0.0871545 |
| SA 1h | 2320.48 | 348.382 | 0.99761 | 0.184912 |
| Me-JA 1h | 1373.37 | 79.5415 | 0.76055 | 0.0651491 |
| Control 6h | 4820.92 | 969.673 | 2.0784 | 0.29366 |
| ABA 6h | 1787.43 | 120.318 | 0.758525 | 0.0438197 |
| ACC 6h | 3841.09 | 1195.67 | 1.37234 | 0.277167 |
| BABA 6h | 5465.09 | 826.935 | 2.16584 | 0.31118 |
| Chitin 6h | 5067.05 | 414.573 | 2.14308 | 0.123742 |
| Epi 6h | 5448.92 | 676.515 | 2.1532 | 0.431055 |
| SA 6h | 4179.56 | 1050.13 | 1.76289 | 0.335479 |
| Me-JA 6h | 2263.77 | 741.449 | 0.87267 | 0.361699 |
Source Transcript PGSC0003DMT400014695 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G18170.1 | +3 | 3e-81 | 260 | 144/210 (69%) | FKBP-like peptidyl-prolyl cis-trans isomerase family protein | chr1:6254291-6255507 FORWARD LENGTH=247 |
