Probe CUST_6668_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6668_PI426222305 | JHI_St_60k_v1 | DMT400036706 | GCCAATAACGACTCGTTGGCGATAATATCTTTGCTTAACATACATAACTATTTGAGGTTT |
All Microarray Probes Designed to Gene DMG400014156
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6602_PI426222305 | JHI_St_60k_v1 | DMT400036705 | CACAAGAGTCATATATGATACAAAACAATCACAAGTTGGATTTGCAAAAGAAACCTGCAG |
| CUST_6668_PI426222305 | JHI_St_60k_v1 | DMT400036706 | GCCAATAACGACTCGTTGGCGATAATATCTTTGCTTAACATACATAACTATTTGAGGTTT |
Microarray Signals from CUST_6668_PI426222305
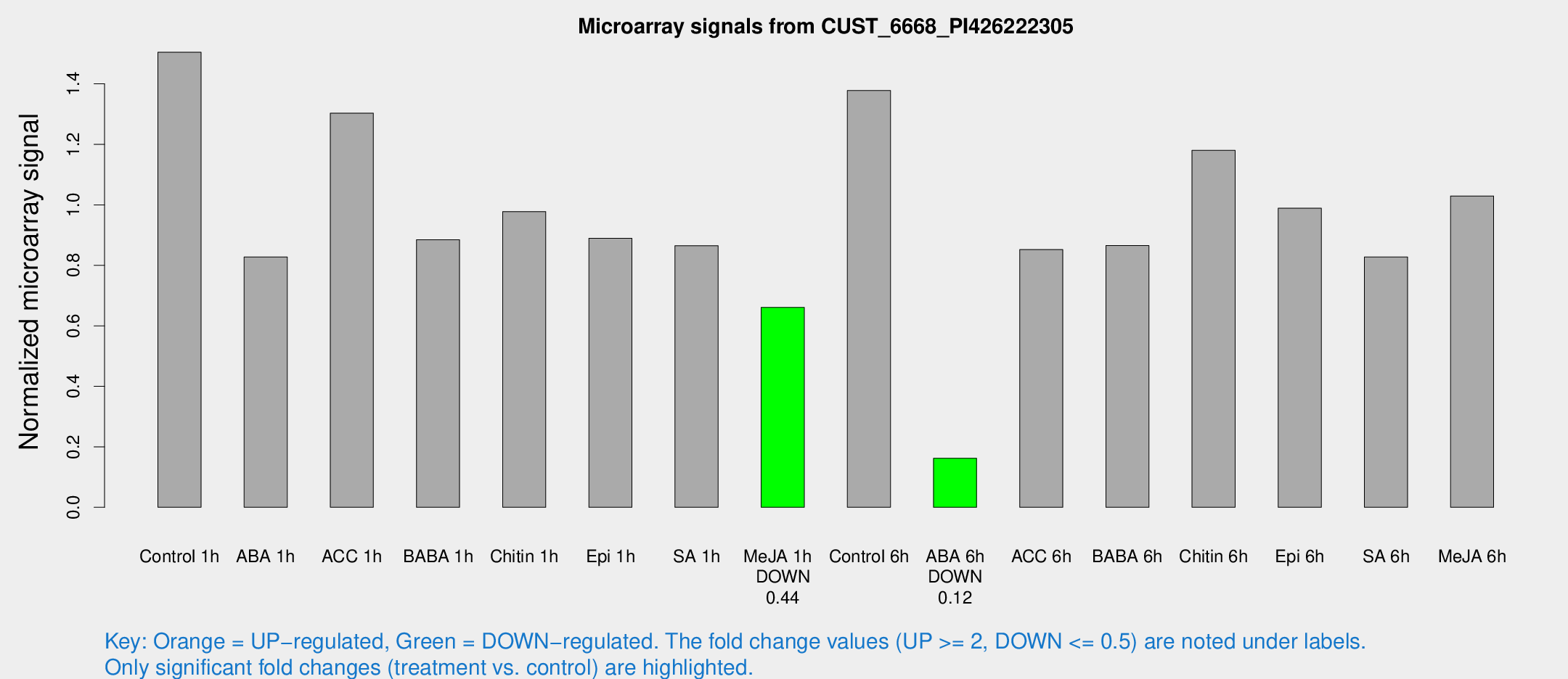
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2897.78 | 249.474 | 1.50461 | 0.0868895 |
| ABA 1h | 1458.19 | 281.091 | 0.827839 | 0.160697 |
| ACC 1h | 2587.18 | 278.655 | 1.30311 | 0.0984575 |
| BABA 1h | 1720.18 | 371.04 | 0.884813 | 0.124533 |
| Chitin 1h | 1699.27 | 173.363 | 0.977845 | 0.0872632 |
| Epi 1h | 1492.11 | 169.209 | 0.88955 | 0.0853196 |
| SA 1h | 1734 | 237.318 | 0.864987 | 0.0948327 |
| Me-JA 1h | 1038.19 | 81.1039 | 0.660779 | 0.0382236 |
| Control 6h | 2792.28 | 594.28 | 1.37802 | 0.211999 |
| ABA 6h | 336.338 | 44.8887 | 0.162199 | 0.0239638 |
| ACC 6h | 1922.76 | 309.338 | 0.852136 | 0.0821871 |
| BABA 6h | 1931.12 | 411.499 | 0.865537 | 0.17511 |
| Chitin 6h | 2408 | 139.412 | 1.18033 | 0.0681771 |
| Epi 6h | 2191.29 | 339.898 | 0.988767 | 0.248056 |
| SA 6h | 1658.75 | 374.228 | 0.827386 | 0.118492 |
| Me-JA 6h | 2019.2 | 325.049 | 1.02932 | 0.103386 |
Source Transcript PGSC0003DMT400036706 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G79720.1 | +2 | 0.0 | 534 | 268/439 (61%) | Eukaryotic aspartyl protease family protein | chr1:29997259-29998951 REVERSE LENGTH=484 |
