Probe CUST_6608_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6608_PI426222305 | JHI_St_60k_v1 | DMT400014657 | TACCGCAGCAAGCTCATACTACGATTTCAACTCGGAATCTGACATTCTCGTGTCATTGCA |
All Microarray Probes Designed to Gene DMG400005725
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6534_PI426222305 | JHI_St_60k_v1 | DMT400014656 | GAAAGCAGTTCAGCTTATTCATCCCCTGCAGCTTCTGCTACGGGGAATAAAAGAAGAAAT |
| CUST_6608_PI426222305 | JHI_St_60k_v1 | DMT400014657 | TACCGCAGCAAGCTCATACTACGATTTCAACTCGGAATCTGACATTCTCGTGTCATTGCA |
Microarray Signals from CUST_6608_PI426222305
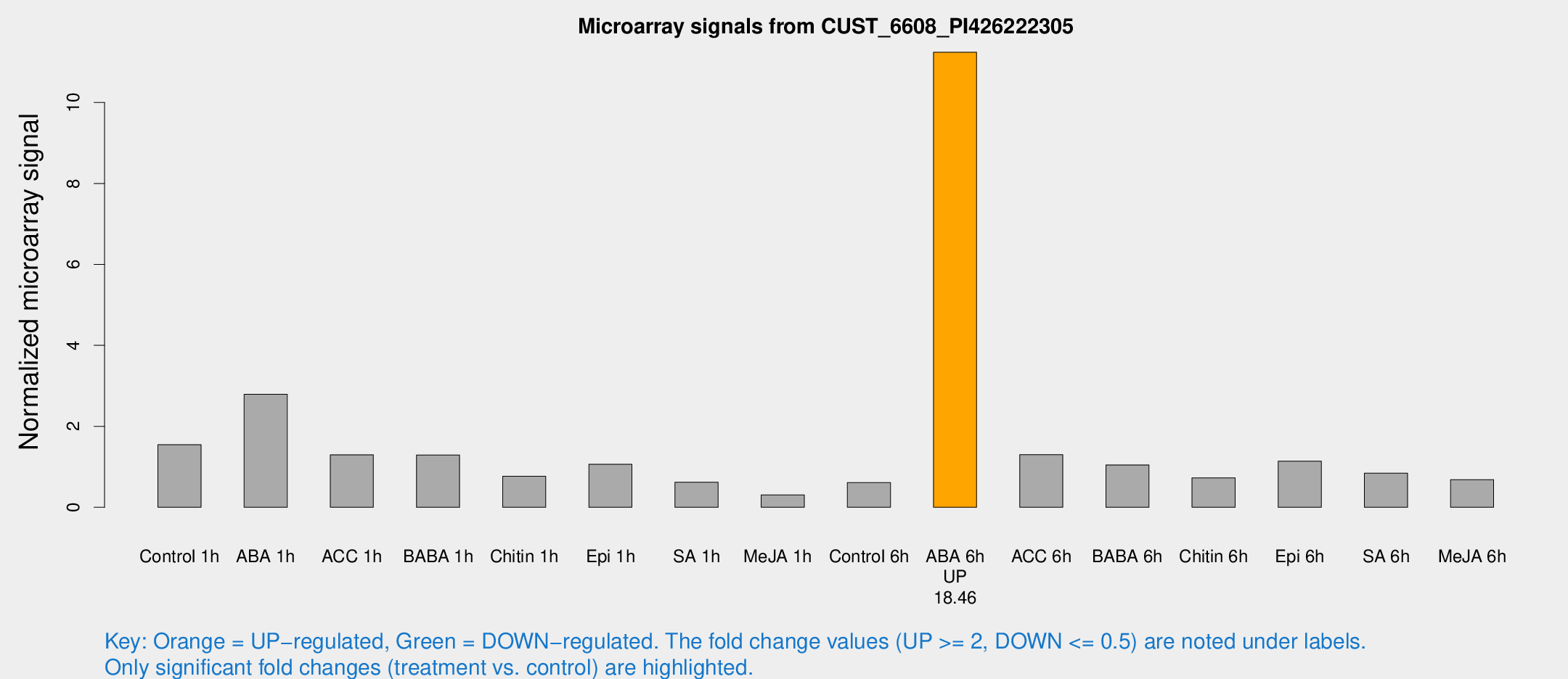
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 563.89 | 112.73 | 1.5462 | 0.380472 |
| ABA 1h | 976.452 | 301.619 | 2.79349 | 1.03288 |
| ACC 1h | 607.157 | 272.949 | 1.29674 | 0.776536 |
| BABA 1h | 471.357 | 126.692 | 1.29181 | 0.276211 |
| Chitin 1h | 272.014 | 85.9049 | 0.766051 | 0.312214 |
| Epi 1h | 395.745 | 145.085 | 1.06233 | 0.65188 |
| SA 1h | 235.801 | 61.9221 | 0.61814 | 0.143344 |
| Me-JA 1h | 92.9617 | 24.4212 | 0.306833 | 0.0655868 |
| Control 6h | 248.876 | 103.851 | 0.608978 | 0.233286 |
| ABA 6h | 4441.36 | 1043.4 | 11.2419 | 2.58087 |
| ACC 6h | 577.652 | 160.638 | 1.30246 | 0.405336 |
| BABA 6h | 435.503 | 114.25 | 1.04743 | 0.237446 |
| Chitin 6h | 273.493 | 36.2858 | 0.725481 | 0.100468 |
| Epi 6h | 486.596 | 133.949 | 1.1406 | 0.265481 |
| SA 6h | 291.457 | 17.6003 | 0.845171 | 0.0913094 |
| Me-JA 6h | 266.088 | 94.7795 | 0.683798 | 0.233411 |
Source Transcript PGSC0003DMT400014657 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G73830.1 | +1 | 3e-11 | 59 | 27/38 (71%) | BR enhanced expression 3 | chr1:27760027-27761346 FORWARD LENGTH=261 |
